
Composition characteristics of the gut microbiota in infants and young children of under 6 years old between Beijing and Japan
Introduction
Microbiota is a microbial community (bacteria, viruses, parasites, fungi) living in a specific environment. In humans, the composition of microbiota depends upon body parts (skin, mouth, vagina, intestines or nostrils) (1). Gut microbiota is the most important one, which is constituted by 1012–1014 microorganisms, representing 2–10 times the number of cells in human body (2). It is now recognized to be beneficial for digestion, metabolism and immunity (3). Therefore, understanding imbalance is an important factor for studying certain diseases, such as chronic inflammatory bowel disease and autoimmune diseases in particular.
Microbiota changes continuously throughout the life, especially in first 6 years after birth. It has been reported that the initial gut microbiota is dependent on the mode of delivery (4). The intestines of naturally-born babies are sown by mother’s vaginal and fecal bacteria, including mainly Bifidobacterium and Lactobacilli (5), while the intestines of the caesarean section-born babies are sown by environmental bacteria, which may not be specific to the intestinal flora (6). Therefore, the gut microbiota changes in the first few years of life until it reaches the compositional characteristics of adult gut microbiota (7).
Intestinal flora has been widely investigated but only one study has been conducted on the infants and young children of two Asian countries (8). So far, one study has investigated the gut microbiota composition in South American populations and four on Asian populations (9). Research focused on the changes in microbial diversity and composition in people with different livelihood patterns, although there is a broad consensus that the pregnancy environment and fetus are sterile before delivery (10). Studies have shown the presence of bacterial DNA in placenta, amniotic fluid and umbilical cord (11,12). Bacterial presence in these compartments can explain its spreading from mother to fetus and non-sterility of the substances ingested during pregnancy (intestine and epithelial cells, placenta, mucus, amniotic fluid, bile and water) (13). Moreover, human enteroviruses are commonly detected in the stool samples of infants. Although the enteroviruses mainly cause mild or asymptomatic infections in healthy children, it can also cause diarrhea (14). The infectious diseases are the cause of many deaths of children, especially among young children living in developing countries/regions, due to poor sanitation, unsanitary drinking water, contaminated food or improper disposal of feces (15).
Therefore, it is important to investigate the differences in the composition of intestinal fecal bacteria/viruses in two Asian countries and to monitor the changes in the intestinal flora at an early stage. In this study, fecal samples were collected from the infants aged 0–6 from Beijing. With the consent of the original author, the data including the original microbiome sequence published by the Japanese CDC in the journal Nature was downloaded. Based on the consistency between this study and the bacterial alternative analysis method issued by the Japanese Centers for Disease Control and Prevention, we compared completely correct data to analyze the composition characteristics of the initial flora of children in different countries and understand the differences and similarities. Then, these data were used (I) to investigate the similarities in the structures and compositions of gut microbiota in infants from Beijing (China) and Japan; (II) to analyze the abundance and structure of fecal bacteria in different ages; (III) to analyze the effects of vaccination, delivery methods (natural or caesarean section) and feeding methods (breastfeeding or formula feeding) on gut microbiota; and (IV) to investigate the similarities in microbe-related signaling pathways the two countries using KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway analyses.
We present the following article in accordance with the MDAR reporting checklist (available at http://dx.doi.org/10.21037/tp-20-376).
Methods
Origin of samples
A total 84 infants, aged 0–6 years, were selected from a community in Tongzhou District, Beijing, China. The study was approved from Ethics Committee of the Seventh Medical Center of Chinese PLA General Hospital (Approval No. 2020-067) and informed consent was signed from the parents of all registered infants. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The biometrics of selected infants, including height, weight, age and bad breadth and other statuses including defecation, feeding, recent vaccination and illness and medication in past two weeks were recorded.
Stool sample collection
The stool samples were collected in sterile specimen boxes. Total bacterial DNA was extracted using QIAamp DNA Stool Mini Kit (QIAGEN, Germany) and stored at −70 °C, which were then sent to Beijing Biomarker Biotechnology Co., Ltd. and Beijing Quantitative Health Co., Ltd. for sequencing and bioinformatics analysis.
Primer for amplification and pre-amplification
Bacterial universal primers were used to amplify 16S rRNA gene V3 + V4 from the stool DNA samples using polymerase chain reaction (PCR). The primer sequences included: bacterial 16S rRNA (V3 + V4) region primers; F-5'-ACTCCTACGGGAGGCAGCA-3', R-5'-GGACTACHVGGGTWTCTAAT-3'; Fungal ITS1 region primer: F-5'-CTTGGTCATTTAGAGGAAGTAA-3', R-5'-GCTGCGTTCTTCATCGATGC-3'; Internal ITS1: F-5'-AACTGCGGAAGGATCATT-3' Internal ITS1: R-5'-GARCCAAGAGATCCRTTG-3'. The conditions of PCR pre-experiment during sample detection were as follows: 95 °C for 5 min (initial denaturation); followed by 30 cycles of 95 °C for one min (denaturation), 50 °C for one min (annealing) and 72 °C for one min (extension); and 72 °C for 7 min (final extension), and then stored at 4 °C for one hour.
Construction of microbial diversity database
PCR purification of target area
The PCR products were screened by mixing with AMPure XP magnetic beads for 5 min at room temperature. The mixture was then placed on magnetic stand for 5 min and supernatant was discarded. The magnetic beads were washed twice with 200 µL of 80% ethanol and the supernatant was discarded each time after 30 sec at room temperature. The magnetic beads were then dried in magnetic strand for 3 min and then re-suspended with 37 µL double distilled water. Then, the suspension was incubated at room temperature for 2 min and placed on magnetic stand for 2 min. 35 µL of supernatant was taken into a new PCR tube and PCR was run with conditions as following: initial denaturation (at 98 °C for 30 sec), followed by 10 cycles of denaturation (98 °C for 10 sec), annealing (65 °C for 30 sec) and elongation (72 °C for 30 sec) and final extension (72 °C for 5 min). The PCR product was run on 1.8% agarose gel for 40 min at 120V.
Magnetic beads purification
The purified and extracted DNA samples and magnetic beads were mixed at a ratio of 1:1.5 in order to screen the DNA fragments and then eluted with 25 µL elution buffer. The fragments were then quantified using NanoDrop quantification method by mixing with Nanodrop 2000 at a mass ratio of 1:1.5. From the 1.8% agarose gel, the DNA fragments were purified using rubber cutting and recycling method.
Sequencing analysis
Illumina HiSeq sequencing platform was used to analyze microbial diversity by paired-end method to construct small fragment libraries for sequencing. The composition of samples was analyzed by splicing and filtering reads, clustering OTUs (Operational Taxonomic Units), and performing species annotation and abundance analysis. The differences in microbial diversity between different samples were further analyzed by alpha diversity and beta diversity.
OTU analysis
OTU is the operation unit of classification. It is an artificially set mark for a certain taxonomic unit (line, species, genus, group, etc.) in order to facilitate analysis in phylogenetic or population genetics studies. According to different similarity levels, all the sequences were divided into OTUs. Generally, if the similarity between sequences is higher than 97%, it can be defined as an OTU, and each OTU corresponds to a representative sequence. UCLUST (version 1.8.0) software was used to cluster tags at a similarity level of 97%, and OTUs were obtained and annotated using Silva (bacteria) and UNITE (fungal) taxonomic databases. Figure 1 shows the number of OTUs of each sample obtained by clustering, where the number of columns shows the number of OTUs of the corresponding sample. Based on the OTU analysis results, taxonomic analysis was performed on the samples at each taxonomic level, and the community structure diagram, species clustering heat map, genera of each sample at the taxonomic level of phylum, class, order, family, genus, and species, taxonomic level phylogenetic tree and taxonomic tree diagram were obtained.
Microbial diversity analysis
Mothur (version v.1.30) software was used to evaluate the Alpha diversity indices of the samples using the standardized number of sequences contained in each sample. The species diversity within each sample was analyzed using Ace, Chao1, Shannon and Simpson indices for each sample at 97% similarity level, and the dilution curve and grade abundance curve of each sample were drawn.
Species differences analysis
Beta diversity analysis was used to compare the differences in species diversity (community composition and structure) of different samples. On the basis of distance matrix, the sample hierarchical clustering (UPGMA) tree, NMDS (Nonmetric Multidimensional Scaling) analysis, sample clustering heat map, sample PCA, PCoA diagram (with grouping information), box plot based on multiple distances, etc. under the corresponding distances were obtained.
The heat map of the samples was drawn using R language tool, where the difference between two samples could be visualized according to the differences in the color gradient. The results of NMDS analysis are shown in Figure 2, where the dots represent samples, different colors represent different groups, and the distance between points represents the degree of difference. Stress value of less than 0.2 indicated reliability of the NMDS analysis.
Gene function prediction analysis
The prediction of gene function and functional gene abundance in the samples were analyzed using 16S functional gene prediction analysis. Based on the Illumina HiSeq sequencing platform, the paired-end sequencing (Paired-End) method is used to construct a small fragment library for sequencing. By splicing and filtering reads, clustering OTUs (Operational Taxonomic Units), and performing species annotation and abundance analysis, the species composition of the sample can be revealed; and further analysis of alpha diversity and beta diversity were conducted.
Statistical analysis
The analysis of significance in differences between groups is mainly used to find biomarkers with statistically significant differences between different groups. According to the biomarker screening criteria [linear discriminant analysis (LDA) score >4], the qualified biomarkers were identified and displayed as icons. The analysis methods included Lefse analysis to screen biomarkers and Metastats analysis that compares the P and q values between two groups at each classification level. P<0.05 indicates that the difference is statistically significant.
Results
Comparison of intestinal flora in age groups
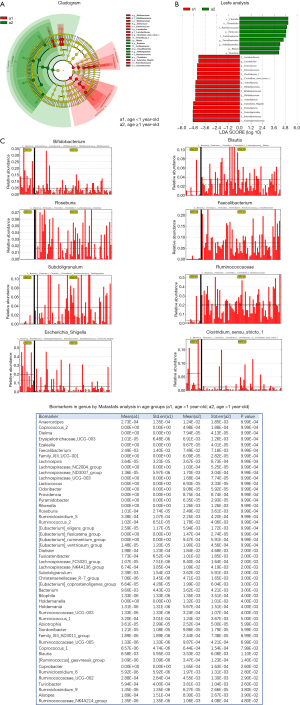
Eighty four healthy infants and young children were divided into two age groups: group A: under 1 year old (11 infants) and group B: 1–6 years old (73 infants). Significant difference was observed between the intestinal floras of both groups. The predominant intestinal flora in group A consisted of mainly Actinomycetes, Bifidobacteria, Lactobacillus, Escherichia, Shigella, Enterobacteriaceae, Proteobacteria, Enterococcus and Clostridium-1, where Clostridium was a narrow case; while the predominant intestinal flora of group B consisted of mainly Firmicutes, Laospirillum, Clostridium, Rumenobacteria, Faetobacter, Blautella, and Rossella. The Lesfe and Matastats analysis showed relatively same abundance of Faebacterium, Anaerostipes, Lachnoclostridium, Blauterella, Rossella, Subdoligranulum, Laevis, Eubacterium eligens, Abiotrophia, Lachnospiraceae_FCS020 and Dialister genera in both groups (Figure 1).
Comparison of intestinal flora in gender-wise groups
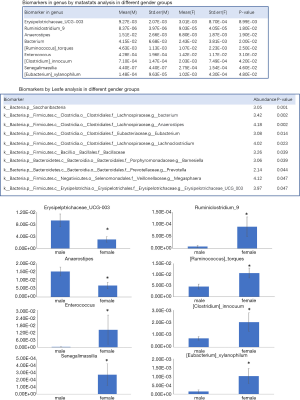
The Lesfe and Matastats analysis in gender-wise groups showed that the abundance of Erysipelotrichaceae_UCG-003 and Anaerostipes was significantly higher in male infants than the female group and young children (P<0.05) (Figure 2).
Comparison of intestinal flora in defecation trait groups
The abundance of intestinal flora in defecation trait groups (abnormal bowel trait group and normal group) showed that Bifidobacterium in the abnormal bowel trait group was significantly more abundant than that in the normal group (P<0.05) (Figure 3). 17.4% (12/69) and 30% (3/15) of the abnormal bowel trait and normal groups used probiotics as daily intervention, respectively, suggesting no statistically significant difference in the defecation trait groups in terms of daily supplement of commercial probiotics. The average ages of the abnormal bowel trait and normal groups were 22.9±10.5 and 22.7±12.7 months, respectively, showing no statistically significant difference. The comparison of defecation regularity showed 26.1% (18/69) of chronic constipation in the normal stool trait group, suggesting that the dry stool and chronic constipation (less than 3 bowel movements per week) had significant positive correlation with Bifidobacterium (P<0.05). This suggested that Bifidobacterium may be involved in the occurrence of chronic constipation.

Comparison of intestinal flora in feeding groups
The feeding groups included breast-feeding group, formula feeding group and mixed feeding group. The abundance of Bifidobacteria and Enterococci in the breast-feeding group was significantly higher than that in the formula feeding and mixed-feeding groups, while Bacteroides and Lacetospirillaceae were lower than the other two groups (Figure 4).
Comparison of intestinal flora in cold /fever and normal groups
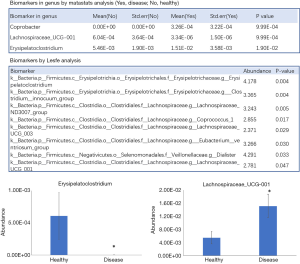
The division on the basis of sickness included two groups: cold/fever group (35/84) and normal group (49/84). In the cold/fever group, 22 cases took medicine after cold and fever (mainly Chinese traditional medicine + antibiotics + antipyretic drugs), while 13 cases did not take medicine (mostly adopting pediatric massage or physical cooling). The differential analysis of the intestinal flora in both groups showed that the abundance of Erysipelatoclostridium in the cold/fever group was significantly higher than that in the normal group (P<0.05), while the abundance of Lachnospiraceae _UCG-001 was significantly lower than that in the normal group (P<0.05) (Figure 5).
Comparison of intestinal flora in halitosis and normal groups
By grouping infants into halitosis and normal groups, the Lesfe and Matastats analysis showed the abundance of Parasutterella and Ruminococcaceae_UCG-003 were significantly higher in the halitosis group than that in the normal group (Figure 6). It has been reported that the abundance of Roseburia, Lachnospiraceae, Alistipes, and Ruminococcaceae in the intestinal flora in HIV infection patients was significantly reduced in comparison with the healthy populations, where the reduction of valeric acid content was positively correlated with the decline in Ruminococcaceae UCG-003 (6).
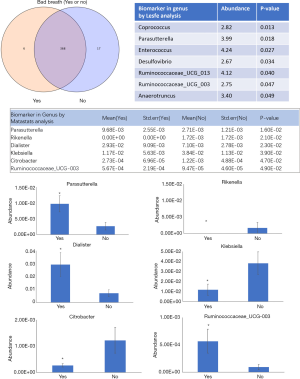
Comparison of intestinal flora in vaccination and normal groups
The infants and young children were divided into vaccination group (10/84) and normal group (74/84). The Lesfe analysis showed that the relative abundance of Ruminococcaceae_NK4A214 in the vaccination group was significantly higher than that in the normal group (Figure 7).
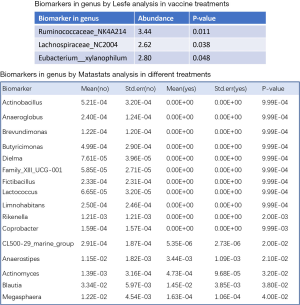
Comparison of intestinal flora of 0-6 year-old infants from Beijing and Japan
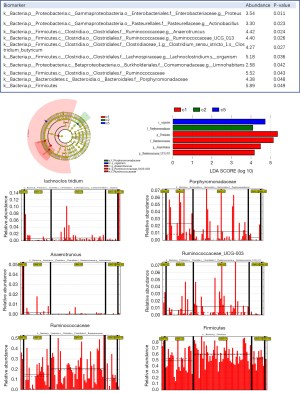
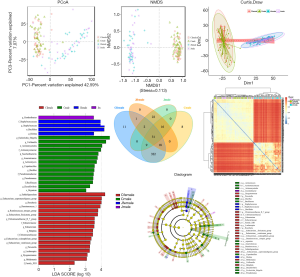
The sequencing data published in Nature by Japanese CDC was downloaded with the consent of the original authors. Based on the consistency of the bacterial population sequencing analysis in Beijing and Japan, the data of both the regions was compared and analyzed. There were significant regional differences in the bacterial flora of infants and children from Beijing and Japan, as well as in bacterial flora distribution on the basis of age and gender. Partial least squares-discriminant analysis (PLS-DA) was used for further investigation of the differences between the two regional groups (Figure 6). The intestinal flora of both regions showed a distinct distribution of microbial species with no crossover. An additional assessment of the PLS-DA analysis, involving the magnitude of the LDA effect, was performed to determine the microbial species which have a greater impact on the differences observed between the groups. As shown in the block diagram in Figure 8, Bacillus, Lactobacillus, Prevotella, Megamonas and Veillonella were significantly increased in 0-6 year-old infants in Beijing as compared with that in Japan. In addition, the abundance of Rumen cocci, Renkenellaceae, and Alipipes strains was significantly reduced (Figure 8).
Discussion
This study systematically analyzed the composition characteristics of intestinal flora in combination with multifactor status from 84 infants and children aged 0–6 years in Beijing, and analyzed the influence of a number of factors on the intestinal flora, such as gender, age, disease status, medication, vaccination status, bowel status and tone status, etc. (16). The bacteria of the genus Prevotella are particularly abundant in African samples (17). Similarly, a study also reported that the bacteria of the genus Prevotella were abundant in samples from adult Baka hunter-gatherers in the Central African Republic, but were not found among American Americans (18). The sequencing data published by Japanese CDC was compared with the sequencing data of this study. The differential analysis suggested that there was unique regional distribution of intestinal flora in infants and young children between Beijing and Japan (19).
In the age groups, significant differences were observed between their intestinal floras. The intestinal bacterial flora in infants of less than 1 year old with higher relative abundance mainly included Actinomycete, Bifidobacterium, Lactobacillus, Escherichia, Shigella, Enterobacteriaceae, Proteobacteria, Enterococcus and Clostridium-1, while those in the infants and children aged 1–6 years old mainly included Firmicutes, Laospirillaceae, Clostridium, Rumenomycetes, Faeculus spp., Blautella spp. and Rossella spp (20) (Figure 1). This suggested a unique distribution of bacterial flora at different ages in infants and children during development. The Lasfe and Matastats analysis of the gender-wise groups demonstrated significantly higher relative abundance of Erysipelotrichaceae_UCG-003 and Anaerostipes in the male group (P<0.05, Figure 2). Some studies pointed out that the relative abundances of Erysipelotrichaceae and Lachnospiraceae were higher in the male and female groups, respectively (21-23). These two fungal families were not detected in other studies. This suggested that infant’s gender might not be a key factor for the colonization of intestinal flora, but it might have a certain impact on the composition of intestinal flora during the growth and development of infants.
At present, few studies have been conducted on the influence of gender on intestinal flora, including those conducted on adults (24). The results found that the difference between the intestinal flora of male and female is affected by the mass index, suggesting an important impact on metabolic diseases and intestinal inflammatory diseases (25). Therefore, the gender is needed to be reconsidered when colonizing intestinal flora in infancy. This study did not found significant effect of gender on the diversity and abundance of the intestinal flora in infants, but found significant differences at the genus level. The relative abundance of the bacterial genera and Enterococcus [Clostridium]innocuum was relatively high in male group, and that of Ruminiclostridium, Eubacterium, Senegalimassilia was relatively high in the female group, where the genus Senegalimassilia was specific to female group only and not detected in the male group. This study only analyzed the differences in intestinal bacteria caused by gender. More studies are needed to study the impact of these differentially dominant bacteria on the short-term and long-term health of infants of different genders, in order to prevent or treat metabolic and other related disorders in adulthood.
Combining Rome V criteria and Bristol stool scale, the stool samples were grouped according to the characteristics of the stool during defecation. Soft strips were considered as normal, and dry, hard or thin and watery stools were considered as abnormal. By grouping, 15/84 of the samples were considered as abnormal stools (13/15 (86.7%) of the samples were dry and hard, 2/15 (13.3%) of the samples were loose), and 69/84 of the samples had normal stool characteristics. Using the differential markers of Lesfe and Matastats, the common genus associated defecation abnormalities were Morganella and Desulfovibrio (26). The relative abundance of Desulfovibrio in the abnormal bowel trait group was significantly lower than that of the normal group (P<0.05). In order to analyze the correlation of changes in defecation traits with flora disorders, further studies are needed to analyze the relevant dominant flora and regulatory mechanism of defecation traits.
In the age groups, the proportion of bifidobacteria in the intestine of infants was higher than that in the young and middle-aged, and was also higher than the proportion of other species of dominant flora in human intestines. The proportion of Lactobacillus mirabilis and Bacteroides and Clostridium in the intestines of young adults was slightly higher than the infants. On the other hand, the proportion of desulfovibrio and Clostridium prasectus in the intestines of middle-aged adults was slightly higher than that of the young adults and significantly higher than infants. In a study, the Bacteroides, Lactobacillus, and Clostridium tenuifolia were reported to be more closely related to each other between the intestines of juvenile group and the longevity group in Changshou Village of Bama, Guangxi, than the middle-aged and elderly groups (27). This study indicated that there are differences in the intestinal flora of healthy people at different physiological ages. Therefore, the selection of age-related donors for fecal microbiota transplant (FMT) may be more reasonable for adolescents. In the infants over one year old and young children, the intestinal flora becomes stable, which is conducive to the body’s metabolism and growth. Therefore, investigation of the distribution of intestinal flora among healthy infants and young children aged 0–6 years old is important.
The proportion of Bifidobacterium and Enterococcus in the breast-feeding group was significantly higher than those of the pure milk-feeding group and the mixed-feeding group, while the proportion of Bacteroides and Lacetospirillaceae was lower in the breast-feeding group than the other two groups.
The abundance of Erysipelatoclostridium bacteria in the epilepsy group was significantly higher than that in the healthy group, suggesting that the febrile seizures and epilepsy, caused by cold and fever in infants and young children, might be similarly correlated to the increase in the bacteria. Therefore, during cold and fever, regardless of taking medicines, the use of feces of infants and young children as a donor for fecal bacteria transplantation is not suitable and may increase the potential risk of brain-gut axis immune endocrine metabolism.
In the diseased and healthy groups, Lachnospiraceae_UCG-001 and Erysipelatoclostridium were bacterial genera found in different abundances. The abundance of Erysipelatoclostridium in the diseased group was significantly higher than that in the healthy group (P<0.05), while that of Lachnospiraceae _UCG-001 was lower than the healthy group (P<0.05). As compared to the colorectal cancer patients, the intestinal flora healthy populations contained mainly clostridia and lachnospiraceae.
In comparison with Japanese children, the intestinal microbiota of children aged 0–6 years old in Beijing is more abundant in Bacteroides. In addition, the genus Prevotella is particularly more abundant in Chinese children’s than Japanese. Several studies have reported genus Prevotella in in the intestinal flora f Asians, but none has reported in from American and European population, despite investigating abundant number of study samples. This study indicated genera Bacteroides and Collinsella in abundance in Chinese population. This study also showed higher relative abundance of genera Bacteroides and Anaerostipes in male group, and that of Ruminiclostridium, Eubacterium and Senegalimassilia in female group. It was consistent with the strain-specific expression results of Japanese children. Studies have reported that the mode of delivery affects the microbial diversity of intestines and their colonization patterns (28). There are bacterial floras in the first three months after birth. However, currently, none of the evidences have shown the effect of delivery mode on the intestinal bacterial flora. The results of this study showed that the intestinal microbiota of infants is only related to the mother’s diet. The bacteria of genera Clostridium and Bifidobacterium in the feces of breast-fed children were lower than those of the non-breast-fed children.
The data comparison of the intestinal flora of infants between Beijing and Japan showed Lactobacillus, Enterococcus, and Bifidobacterium to be the dominant bacteria in the intestines of infants under 1 year old in both regions. Among them, the proportion of Bifidobacterium, Bacillus and Eubacterium rectum in infants aged 0–6 in Japan was significantly higher than that in Beijing. The proportion of Bifidobacteria in infants under 1 year old in Japan was significantly higher than that in the infants over 1 year old in the same region. Eubacteria rectum was the dominant species in Japanese infants of over 1 year old. In Japan, the proportion of Staphylococcus in infants under 1 year old was significantly higher than that in the other groups; while the proportion of intestinal Lactobacillus and Enterobacter in infants under 1 year old in Beijing was significantly higher than that in the other groups. The proportions of Laospirillum, Clostridium prastilus, Eubacteria fecalsterol, and rare Micrococcus in the infants over 1 year old in Beijing area were significantly higher than those of other groups (P<0.001). As shown in the block diagram, as compared to 0–6 year-old infant group in Japan, Beijing’s 0–6 year-old infant group has significantly more abundance of Bacillus, Lactobacillus, Prevotella, Macrophages and Weillonella Up, in their intestinal flora.
The enrichment analysis of 100 core floras in Chinese and Japanese children identified 43 significantly enriched KEGG signaling pathways, including inflammation pathways, PI3K-Akt (phosphatidylinositol 3 kinase-protein kinase B) signaling pathways, AGE-RAGE signaling pathways in genetic diseases, congenital herpes virus infection, fluid shear stress, human cytomegalovirus infection, IL-17 signaling pathway, neuro-active ligand-receptor interaction, hepatitis B, MAPK signaling pathway, metabolic pathway, TNF signaling pathway, human papillomavirus infection, measles signaling pathway and human T-cell leukemia virus. These results showed that Chinese and Japanese children might adopt PI3K-Akt signaling pathway, MAPK signaling pathway, and TNF signaling pathway using multiple targets, directly or indirectly affecting children’s health.
Taking one year old as boundary, there were significant differences in the intestinal micro-ecology of children between infants under one year of age group and children of 1–6 years old group in Beijing. Based on the age and gender of the bacteria, there were also obvious regional differences in the intestinal flora of infants between Beijing and Japan in terms of feeding methods and the incidence of children. Through comparison, the dominant flora of children aged 0–6 years old can be obtained, but the in-depth discussion and commercial development of its mechanism can be used on the premise that it is safe and feasible and can replace fecal bacteria transplantation.
Acknowledgments
Funding: This study was supported by the Wu Jieping Medical Foundation [320.6750.18172].
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at http://dx.doi.org/10.21037/tp-20-376
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tp-20-376
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tp-20-376). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved from Ethics Committee of the Seventh Medical Center of Chinese PLA General Hospital (Approval No. 2020-067) and informed consent was signed from the parents of all registered infants.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Cherniavska YI, Pokhylko VI, Znamenska TK, et al. Effect of enos gene polymorphism on the course of early onset bacterial infections in premature infants. Wiad Lek 2020;73:1237-40. [Crossref] [PubMed]
- Pelkonen T, Urtti S, Dos Anjos E, et al. Aetiology of bacterial meningitis in infants aged <90 days: Prospective surveillance in Luanda, Angola. Int J Infect Dis 2020;97:251-7. [Crossref] [PubMed]
- Awasthi S, Kesarwani N, Verma RK, et al. Identification and management of young infants with possible serious bacterial infection where referral was not feasible in rural Lucknow district of Uttar Pradesh, India: An implementation research. PLoS One 2020;15:e0234212 [Crossref] [PubMed]
- Orman G, Kukreja MK, Vallejo JG, et al. Accuracy of MR Imaging for Detection of Sensorineural Hearing Loss in Infants with Bacterial Meningitis. AJNR Am J Neuroradiol 2020;41:1081-6. [Crossref] [PubMed]
- Rahman AE, Herrera S, Rubayet S, et al. Managing possible serious bacterial infection of young infants where referral is not possible: Lessons from the early implementation experience in Kushtia District learning laboratory, Bangladesh. PLoS One 2020;15:e0232675 [Crossref] [PubMed]
- Brower LH, Wilson PM, Murtagh-Kurowski E, et al. Evaluation for Neonatal HSV in Infants Undergoing Workup for Serious Bacterial Infection: A 5-Year Retrospective Review. Hosp Pediatr 2020;10:463-70. [Crossref] [PubMed]
- Zwittink RD, van Zoeren-Grobben D, Renes IB, et al. Dynamics of the bacterial gut microbiota in preterm and term infants after intravenous amoxicillin/ceftazidime treatment. BMC Pediatr 2020;20:195. [Crossref] [PubMed]
- Applegate JA, Ahmed S, Harrison M, et al. Caregiver acceptability of the guidelines for managing young infants with possible serious bacterial infections (PSBI) in primary care facilities in rural Bangladesh. PLoS One 2020;15:e0231490 [Crossref] [PubMed]
- Guenther T, Mopiwa G, Nsona H, et al. Feasibility of implementing the World Health Organization case management guideline for possible serious bacterial infection among young infants in Ntcheu district, Malawi. PLoS One 2020;15:e0229248 [Crossref] [PubMed]
- Wammanda RD, Adamu SA, Joshua HD, et al. Implementation of the WHO guideline on treatment of young infants with signs of possible serious bacterial infection when hospital referral is not feasible in rural Zaria, Nigeria: Challenges and solutions. PLoS One 2020;15:e0228718 [Crossref] [PubMed]
- Ramgopal S, Wilson PM. Automated Versus Manual Band Counts for the Diagnosis of Invasive Bacterial Infections in Infants Who Are Febrile. J Pediatr 2020;221:246-250.e3. [Crossref] [PubMed]
- Gordon SM, Srinivasan L, Taylor DM, et al. Derivation of a metabolic signature associated with bacterial meningitis in infants. Pediatr Res 2020;88:184-91. [Crossref] [PubMed]
- Roßberg S, Keller T, Icke K, et al. Orally applied bacterial lysate in infants at risk for atopy does not prevent atopic dermatitis, allergic rhinitis, asthma or allergic sensitization at school age: Follow-up of a randomized trial. Allergy 2020;75:2020-5. [Crossref] [PubMed]
- Klouda TM, Wang H, Yaeger JP. Association of Cough Status With Bacterial Infections in Febrile Infants. Hosp Pediatr 2020;10:185-9. [Crossref] [PubMed]
- Raba AA, Krebit I. Definite bacterial infection in recently vaccinated febrile infants. J Paediatr Child Health 2020;56:889-92. [Crossref] [PubMed]
- Applegate JA, Ahmed S, Khan MA, et al. Early implementation of guidelines for managing young infants with possible serious bacterial infection in Bangladesh. BMJ Glob Health 2019;4:e001643 [Crossref] [PubMed]
- Akagawa Y, Kimata T, Akagawa S, et al. Optimal bacterial colony counts for the diagnosis of upper urinary tract infections in infants. Clin Exp Nephrol 2020;24:253-8. [Crossref] [PubMed]
- Fleischer E, Neuman MI, Wang ME, et al. Cerebrospinal Fluid Profiles of Infants ≤60 Days of Age With Bacterial Meningitis. Hosp Pediatr 2019;9:979-82. [Crossref] [PubMed]
- Barry HC. Laboratory-Based Prediction Model Can Rule Out Serious Bacterial Infections in Febrile Infants. Am Fam Physician 2019;100:440. [PubMed]
- Bonilla L, Gomez B, Pintos C, et al. Prevalence of Bacterial Infection in Febrile Infant 61-90 Days Old Compared With Younger Infants. Pediatr Infect Dis J 2019;38:1163-7. [Crossref] [PubMed]
- Huang YH, Yan JH, Kuo KC, et al. Early antibiotics use in young infants with invasive bacterial infection visiting emergency department, a single medical center’s experience. Pediatr Neonatol 2020;61:155-9. [Crossref] [PubMed]
- Poletto E, Zanetto L, Velasco R, et al. Bacterial meningitis in febrile young infants acutely assessed for presumed urinary tract infection: a systematic review. Eur J Pediatr 2019;178:1577-87. [Crossref] [PubMed]
- Davis J, Lehman E. Fever Characteristics and Risk of Serious Bacterial Infection in Febrile Infants. J Emerg Med 2019;57:306-13. [Crossref] [PubMed]
- Brown BP, Jaspan HB. Compositional analyses reveal correlations between taxon-level gut bacterial abundance and peripheral T cell marker expression in African infants. Gut Microbes 2020;11:237-44. [Crossref] [PubMed]
- Sobczak A, Klepacka J, Amrom D, et al. Umbilical catheters as vectors for generalized bacterial infection in premature infants regardless of antibiotic use. J Med Microbiol 2019;68:1306-13. [Crossref] [PubMed]
- Ramchandar N, Gierhart S, Creppage KE, et al. Epidemiology of Serious Bacterial Infections in Infants Less Than 90 Days in a Military Health System Cohort. Pediatr Infect Dis J 2019;38:849-53. [Crossref] [PubMed]
- Pokhylko V, Kovalova O, Cherniavska Y, et al. Development of arterial hypotension in premature infants with early onset bacterial infections: tools of clinical predication. Wiad Lek 2019;72:1068-73. [Crossref] [PubMed]
- Chi C, Xue Y, Lv N, et al. Longitudinal Gut Bacterial Colonization and Its Influencing Factors of Low Birth Weight Infants During the First 3 Months of Life. Front Microbiol 2019;10:1105. [Crossref] [PubMed]


