
An interesting Mybpc3 heterozygous mutation associated with bicuspid aortic valve
Introduction
Aortic stenosis (AS) is a common heart valvular disease characterized by progressive valve obstruction and left ventricular remodeling (1,2). It’s morbidity in elderlies of the western world amounted to 12.4% (3), morbidity of AS in juveniles has yet to be reported. It is well known that AS is mainly caused by rheumatic fever sequelae, congenital aortic valve dysplasia or senile aortic valve calcification. Patients can be asymptomatic during the compensatory period, and most patients with severe stenosis suffer from burnout, dyspnea, angina, dizziness or syncope, and in some cases sudden death. Valvular stenosis causes pressure overload of the left ventricle, which in turn triggers a prolonged hypertrophic response. Aortic valve replacement should be considered over time as the patient transition from hypertrophy to heart failure (4).
Bicuspid aortic valve (BAV) is a common congenital heart defect affecting up to 2% of adults today, and potentially the main causes of AS (5). BAV complications contain aortic regurgitation (13–30%), infective endocarditis (2–5%), AS (12–37%), and ascending thoracic aorta dilatation (20–50%) (6). BAV is autosomal dominant with incomplete penetrance and male predominance (7). Myosin binding protein C (Mybpc3) gene sequence and its protein structure were first reported back in 1997 (8). Mybpc3 contains more than 21,000 bps and 34 coding exomes (9). It is well-recognized mutations in specific functional domains or protein translation modification sites can alter protein conformation, protein–ligand binding, or protein–protein interaction (10). Mybpc3 is a member of the intracellular immunoglobulin superfamily, including 11 domains. The current structure and function of Mybpc3 are not fully understood. According to the UniProtKB database, seven conserved domains are I-set (pfam07679), IG (smart00409), IG_like (smart00410), Ig (c11960), I-set (pfam07679), FN3 (cl00065), and Ig (cl11960). The relationship between the location of Mybpc3 mutation and AS/BAV may provide a novel perspective of understanding this disorder.
In this study, we studied a BAV family with two affected and one unaffected member. Whole exome sequencing (WES) was utilized to identify possible disease-causing genes or variants. Paired end reading was aligned with the GRCh37/hg19 human reference sequence. Through comprehensive Clinvar and Genome Analysis Toolkit (GATK) analyzing, Binary Alignment/Map (BAM) and Variant Call Format (VCF) files were produced. An interesting Mybpc3 gene is heterozygous mutation (Ala58Val) was identified in an 11-year-old female proband with BAV. The proband’s parents were also subjected to WES in order to confirm possible disease-causing genes or variants. Our results indicate that Mybpc3 mutation may be associated with BAV and should be screened in prospective clinical practice to encourage early intervention. We present the following article in accordance with the MDAR reporting checklist. Available at http://dx.doi.org/10.21037/tp-20-81.
Methods
Clinical presentation
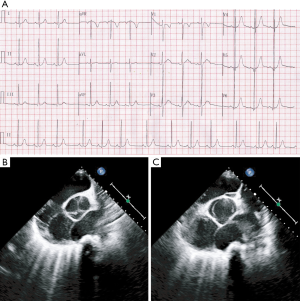
An 11-year-old female proband was admitted for 8 months of continuous precordial discomfort. No sign or symptom was identified during physical examinations. A 12-lead electrocardiogram (ECG) in resting time showed sinus arrhythmia, and minor hypertrophy in the left ventricular (RV5 =42 mm) (Figure 1A). Biochemical metabolism, myocardial enzyme, and cardiac computed tomography angiography were also performed and showed normal results. Further ultrasound cardiogram indicated and recognized that proband suffered from symptoms of BAV (Figure 1B,C). Proband’s father was diagnosed with AS. Patients who have BAV should be monitored regularly to prevent infective endocarditis. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013), and was approved by the Ethics Committee of Children’s Hospital of Shanghai Jiaotong University (No. 2019R002-F01). The patients’ parents provided written informed consent for publication.
WES
DNA libraries of constructs and WES assay were carried out according to the manufacturer’s instructions. Briefly, a whole-blood genomic DNA extraction kit (Tiangen, China) was utilized to isolate genomic DNA, 1 µg DNA was used for WES assay. Specific experimental procedures and experimental instruments are detailed in previous publications (11).
Sanger sequencing and data analysis
Mybpc3 mutation was confirmed via Sanger sequencing. Primers were designed to cover the known mutation sequence. Forward primer: 5'- CGGGCAGGAGTGAAGGTG-3' and reverse primer: 5'- GCCACAGCAAAGGCAAGAAA-3'. PCR products were resolved and purified using QIAquick kit (Qiagen, USA), Sanger sequencing was carried out at Suzhou Hong Xun Biotechnology Co., Ltd.
Realtime-PCR (RT-PCR) was utilized to detect expression of Mybpc3 mRNA, 200 µL of whole-blood sample was used to extract RNA according to protocol (PrimeScript™ RT Master Mix, takara). Primers were designed before and after the mutation to explore whether the mutation altered its expression. Before the mutation forward primer: 5'- GGGGAAGAAGCCAGTCTCAG-3' and reverse primer: 5'- CAGGCCGTACTTGTTGCTG-3'. After the mutation forward primer: 5'- TCAAGCTCAGCAGCTCTCAA-3' and reverse primer: 5'- CATTTGCCCTTGAACCACTT-3'.
The data were filtered and analyzed in our previous study (11). Briefly, BWA-0.710 software was utilized to compare with human genome database (GRCh 37/hg 19). Then promising data were filtered and further compared with the 1,000 Genomes Project, Exome Variant Server, Exome Aggregation Consortium databases, gnomAD, Human Gene Mutation Database (HGMD), Clinvar, and Online Mendelian Inheritance in Man (OMIM). A series of prediction software (Sorts intolerant from tolerant, Polyphen-2, Genomic evolutionary rate profiling, and Mutation Taster) were utilized to predict mutation effects.
Results
General mutation characteristics
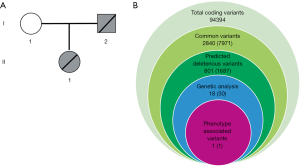
As Figure 2A,B showed, 94,394 variants were detected and these variants were further annotated and filtered by Ingenuity Variant Analysis. A total of 91,554 common variants were filtered and eliminated taken into account their frequencies (Minor Allele Frequency <0.05) according to standards of Exome Aggregation Consortium, 1,000 Genomes Project, Exome Sequencing Project, or gnomAD. Thirty variants in 18 genes were determined and identified through analysis. Finally, the Mybpc3 mutation was detected and selected after rigorous analysis linking Mybpc3 to BAV phenotype (Figure 2B). These rare phenotype-related variants are now classified following the American College of Medical Genetics and Genomics/Association for Molecular Pathology guidelines.
A heterozygous mutation of Mybpc3 gene (Ala58Val) was first identified in the BAV pedigree from WES results. The mutation site c.173C>T is located in exome two of Mybcp3. The proband and her father were confirmed to be heterozygous carriers of 173 C>T hybridization, and her mother was homozygous negative of the mutation as showed through Sanger sequencing (Figure 3A,B). The expression of Mybpc3 mRNA in the proband and her father was reduced in comparison to proband’s mother. Furthermore, RT-PCR results exhibited that relative Mybpc3 mRNA levels in the proband and her father were almost half of proband’s mother (Figure 3C). This result also indicates that p. Ala58Val alter Mybpc3 expression, may play crucial roles for heritable BAV. Importantly, this mutation (c.173C>T) is located in the conserved region of Mybpc3 (Figure 3D), indicating this mutation may affect Mybpc3’s protein function. Western blot results also exhibited that relative Mybpc3 protein levels in the proband and her father were almost half of proband’s mother (Figure 3E).
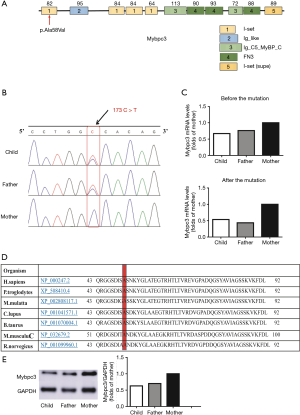
Mutation analysis
In order to further explore the possibility of Mybpc3 mutation (p.Ala58Val) having functional effects of the diseases occurrence, the structure of mutant Mybpc3 protein was compared to a wild type of Mybpc3 (Figure 4A,B). In this study, Mybpc3 heterozygous variant p.Ala58Val was located in the I-set domain (Figure 4C). Light blue indicates myosin. Light green indicates F-actin, C0 domain and C1–C2 domain participate in the interaction of S2 and S1 structures of myosin, forming muscle contraction, and C0 domain is also responsible for in combination with F-actin (Figure 4D). According to prediction, the mutation site plays a critical role in the proper folding of Mybpc3 protein structure. The probable mutation effect may be one of the possible causes leading for heritable BAV.
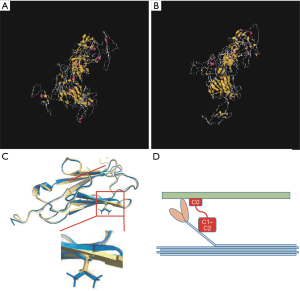
Discussion
In this study, clinical phenotype and genotype of a mutated BAV pedigree were collected and analyzed to investigate a potential disease-causing variant. Three interesting findings are as follows: (I) Ala58Val heterozygous mutation of Mybpc3 is first reported in this BAV pedigree; (II) Protein modeling assay indicates that this mutation could reduce Mybpc3 protein stability; (III) The proband and her father’s Mybpc3 mRNA and protein levels in whole blood were only half of proband’s mother. Indicating that the Mybpc3 (p.Ala58Val) mutation affects Mybpc3 expression, which may play a crucial role for heritable BAV.
It is well recognized that extensive heterogeneity of BAV were due to a combination of genetic, and hemodynamic factors that serve as phenotype modulators. Nine possible BAV genes (NOTCH1, EGFR, AXIN1, GATA5, ENG, PDIA2, NOS3, TGFBR2, and NKX2-5) were employed in forty-eight BAV patients to detect pathogenic variants in AXIN1 (12-14), ENG (13), GATA5 (15-18), NOTCH1 (16-18), and PDIA2 (13). In summary, within the known non-syndromic BAV cases, the incidence of causative gene mutations is less than 1%, which hinders understanding of the disease mechanism and the development of treatment strategies.
In order to further explore the possibility of the Mybpc3 mutation (p.Ala58Val) functional effects of the diseases occurrence, we compared mutant Mybpc3 structure to a wild type copy of Mybpc3 (Figure 4A,B). Mybpc3 heterozygous variant p.Ala58Val is located in the I-set domain and Mybpc3 is a filament protein playing a critical role in modulating muscle contraction. Mybpc3 mutation is the second leading cause of hypertrophic cardiomyopathy (HCM). Mybpc3 binds to myosin at two binding sites, one at its C-terminus and another at its N-terminus. The N-terminal binding site is composed of immunoglobulin domains C1 and C2, connected by flexible ligands and interacting with the myosin S2 by phosphorylation regulation (19). Through GPS database we found the lack of phosphorylation, methylation, or sumoylation of mutated Mybpc3 (p.Ala58Val). Further analysis on the consequence of Mybpc3 heterozygous variant revealed that it may disrupt C0 domain, playing critical roles in combining with myosin and F-actin. As we all know that myosin and F-actin having vital roles in myocardium, posttranslational modification plays a major role in disease progression, the Mybpc3 heterozygous variant may be one of the probable reasons for this BAV family.
In reality, lacking in experimental validation is the main focus of this research. However, a systematic bioinformatics analysis was able to demonstrate the pathogenicity and functionality of the impact of mutation. Furthermore, RT-PCR assay was utilized to analyze Mybpc3 expression among the proband and her parents. The Mybpc3 mRNA expression in the proband and her father was only half of proband’s mother (Figure 3C); Western blot results also exhibited that relative Mybpc3 protein levels in the proband and her father were almost half of proband’s mother (Figure 3E), indicating that p.Ala58Val alters Mybpc3 expression. Our results report an interesting mutation in Mybpc3 gene mutation that is in association with BAV, and the heterozygosity mutation site (Ala58Val) was never reported previously. The p.Ala58Val mutation was first detected in the BAV proband and her father. This mutation is in the conserved area of Mybpc3 protein, and located in I-set domain. Protein modeling assay indicates that the mutation could reduce the hydrophobicity of Mybpc3 protein and its stability.
More than 350 individual Mybpc3 gene mutations have been observed to be related with HCM, making it one of the most frequently mutated genes in HCM (20,21). Studies have shown that Mybpc3 may, through the combination of its N-terminus and myosin Subparticulate-part 2, reduce actin ATPase activity; its phosphorylation could reverse this process. The mechanism of Mybpc3 mutation may be involved in the termination of the phosphorylation process (22). In this study, Mybpc3 heterozygous variant p.Ala58Val was located in the I-set domain of Mybpc3, and the mutation site plays a critical role in Mybpc3 protein structure (Figure 4C,D). Mark Pfuhl’s team hypothesized that Mybpc3 functioned as a tether which fixed S1 heads in a stationary position, and phosphorylation released S1 heads into an active state (19). The probable mutation effect may be one of the possible causes for this heritable BAV. As Mybpc3 plays a crucial role in HCM, may lead to leaving ventricular hypertrophy. If the mutation decreases Mybpc3 stability, it may disable ATP production or APTase activity or phosphorylation regulation, which could partially explain the phenotypes in our patient. Loss of Mybpc3 phosphorylation may cause a primary increase in calcium sensitivity (23). An increase in calcium transients may also have effects on calcium dependent enzymes such as calcineurin, calmodulin dependent kinase, and protein kinase C all of which have been shown to be important for the initiation of myocardial hypertrophy (24). Many Idiopathic dilated cardiomyopathy (DCM) probands also have congenital defects, including two with BAV with aortic regurgitation. Family history and genetic information have potential roles on individuals with aortic regurgitation. Rare variants in the MYBPC3 gene have been reported in several cases of DCM (25). However, the precise molecular mechanism of how down-regulated Mybpc3 expression affects this system needs further explore.
Conclusions
In conclusion, genetic diagnosis of BAV before the onset of symptoms is crucial. The patient and her father were both confirmed with Mybpc3 heterozygous mutation (Ala58Val), indicting Mybpc3 mutation may be the disease-causing variant for heritable BAV. Systematic analysis not only improves our understanding of this disease etiology, but also contributes to clinical and prenatal diagnosis. Determining the genetic origins of BAV is essential to improve the clinical care of patients as well as to develop tailored therapeutic strategies for monitoring disease progression and preventing related aortopathy.
Acknowledgments
Funding: This work is supported by Shanghai Jiaotong University medical technology crossing project (ZH2018QNA31, ZH2018ZDA26), Shanghai science and technology committee (18411965800, 19411963600), Shanghai Healthy Committee (ZY (2018-2020)-FWTX-3023), research fund for National Natural Science Foundation of China (NSFC) (no. 81900437), Shanghai Children’s hospital (2019YQ06, 2018YQN006) and Shanghai key clinical specialty project(shslczdzk05705). No benefits in any form have been or will be received from a commercial organization directly or indirectly.
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at http://dx.doi.org/10.21037/tp-20-81
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tp-20-81
Peer Review File: Available at http://dx.doi.org/10.21037/tp-20-81
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tp-20-81). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013), and was approved by the Ethics Committee of Children’s Hospital of Shanghai Jiaotong University (No. 2019R002-F01). The patients’ parents provided written informed consent for publication.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Dweck MR, Boon NA, Newby DE. Calcific aortic stenosis: a disease of the valve and the myocardium. J Am Coll Cardiol 2012;60:1854-63. [Crossref] [PubMed]
- Iung B, Vahanian A. Epidemiology of valvular heart disease in the adult. Nat Rev Cardiol 2011;8:162-72. [Crossref] [PubMed]
- Osnabrugge RL, Mylotte D, Head SJ, et al. Aortic stenosis in the elderly: disease prevalence and number of candidates for transcatheter aortic valve replacement: a meta-analysis and modeling study. J Am Coll Cardiol 2013;62:1002-12. [Crossref] [PubMed]
- Bing R, Cavalcante JL, Everett RJ, et al. Imaging and Impact of Myocardial Fibrosis in Aortic Stenosis. JACC Cardiovasc Imaging 2019;12:283-96. [Crossref] [PubMed]
- Prakash SK, Bosse Y, Muehlschlegel JD, et al. A roadmap to investigate the genetic basis of bicuspid aortic valve and its complications: insights from the International BAVCon (Bicuspid Aortic Valve Consortium). J Am Coll Cardiol 2014;64:832-9. [Crossref] [PubMed]
- Della Corte A, Bancone C, Buonocore M, et al. Pattern of ascending aortic dimensions predicts the growth rate of the aorta in patients with bicuspid aortic valve. JACC Cardiovasc Imaging 2013;6:1301-10. [Crossref] [PubMed]
- Gould RA, Aziz H, Woods CE, et al. ROBO4 variants predispose individuals to bicuspid aortic valve and thoracic aortic aneurysm. Nat Genet 2019;51:42-50. [Crossref] [PubMed]
- Carrier L, Bonne G, Bahrend E, et al. Organization and sequence of human cardiac myosin binding protein C gene (MYBPC3) and identification of mutations predicted to produce truncated proteins in familial hypertrophic cardiomyopathy. Circ Res 1997;80:427-34. [Crossref] [PubMed]
- Carrier L, Mearini G, Stathopoulou K, et al. Cardiac myosin-binding protein C (MYBPC3) in cardiac pathophysiology. Gene 2015;573:188-97. [Crossref] [PubMed]
- Wu W, Lu L, Xu W, et al. Whole Exome Sequencing Identifies a Novel Pathogenic RET Variant in Hirschsprung Disease. Front Genet 2019;9:752. [Crossref] [PubMed]
- Xie L, Hou C, Jiang X, et al. A compound heterozygosity of Tecrl gene confirmed in a catecholaminergic polymorphic ventricular tachycardia family. Eur J Med Genet 2019;62:103631. [Crossref] [PubMed]
- Armstrong EJ, Bischoff J. Heart valve development: endothelial cell signaling and differentiation. Circ Res 2004;95:459-70. [Crossref] [PubMed]
- Wooten EC, Iyer LK, Montefusco MC, et al. Application of gene network analysis techniques identifies AXIN1/PDIA2 and endoglin haplotypes associated with bicuspid aortic valve. PLoS One 2010;5:e8830. [Crossref] [PubMed]
- Dargis N, Lamontagne M, Gaudreault N, et al. Identification of Gender-Specific Genetic Variants in Patients With Bicuspid Aortic Valve. Am J Cardiol 2016;117:420-6. [Crossref] [PubMed]
- German Z, Chambliss KL, Pace MC, et al. Molecular basis of cell-specific endothelial nitric-oxide synthase expression in airway epithelium. J Biol Chem 2000;275:8183-9. [Crossref] [PubMed]
- Padang R, Bagnall RD, Richmond DR, et al. Rare non-synonymous variations in the transcriptional activation domains of GATA5 in bicuspid aortic valve disease. J Mol Cell Cardiol 2012;53:277-81. [Crossref] [PubMed]
- Bonachea EM, Chang SW, Zender G, et al. Rare GATA5 sequence variants identified in individuals with bicuspid aortic valve. Pediatr Res 2014;76:211-6. [Crossref] [PubMed]
- Shi LM, Tao JW, Qiu XB, et al. GATA5 loss-of-function mutations associated with congenital bicuspid aortic valve. Int J Mol Med 2014;33:1219-26. [Crossref] [PubMed]
- Ababou A, Rostkova E, Mistry S, et al. Myosin binding protein C positioned to play a key role in regulation of muscle contraction: structure and interactions of domain C1. J Mol Biol 2008;384:615-30. [Crossref] [PubMed]
- Behrens-Gawlik V, Mearini G, Gedicke-Hornung C, et al. MYBPC3 in hypertrophic cardiomyopathy: from mutation identification to RNA-based correction. Pflugers Arch 2014;466:215-23. [Crossref] [PubMed]
- Schlossarek S, Mearini G, Carrier L. Cardiac myosin-binding protein C in hypertrophic cardiomyopathy: mechanisms and therapeutic opportunities. J MOL CELL CARDIOL 2011;50:613-20. [Crossref] [PubMed]
- Flashman E, Redwood C, Moolman-Smook J, et al. Cardiac myosin binding protein C: its role in physiology and disease. CIRC RES 2004;94:1279-89. [Crossref] [PubMed]
- Knöll R. Myosin binding protein C: implications for signal-transduction. Journal of muscle research and cell motility 2012;33:31-42. [Crossref] [PubMed]
- Kubis HP, Hanke N, Scheibe RJ, et al. Ca2+ transients activate calcineurin/NFATc1 and initiate fast-to-slow transformation in a primary skeletal muscle culture. Am J Physiol Cell Physiol 2003;285:C56-63. [Crossref] [PubMed]
- Wells QS, Ausborn NL, Funke BH, et al. Familial dilated cardiomyopathy associated with congenital defects in the setting of a novel VCL mutation (Lys815Arg) in conjunction with a known MYPBC3 variant. Cardiogenetics 2011;1:e10. [Crossref] [PubMed]


