Expression of Wilms tumor gene in high risk neuroblastoma: complementary marker to tyrosine hydroxylase for detection of minimal residual disease
Introduction
Neuroblastoma (NB) is derived from primordial neural crest cells that eventually populate the adrenal medulla, sympathetic ganglia, and other sites (1). It is the most common extracranial solid tumor in children under 5 years of age (2-4) and is characterized by clinical heterogeneity. Around 40% to 50% of NB patients in advanced stage 4 presents with dissemination in bone marrow (BM), bone, distant lymph nodes, liver and other organs (4,5). While NB can spontaneously mature (6-9), this enigmatic tumor more often presents with metastatic disease at diagnosis and it is this aggressive propensity which places it among the deadliest pediatric tumors despite intensive multimodal therapy including stem cell transplant (1,2,10,11). Though histologically NB is classified under small round cell tumors (SRCTs) they can be ambiguous. Thus, it is essential to ascertain reliable tumor markers by molecular biological and/or cytogenetic analyses to facilitate appropriate diagnosis and follow-up including the detection of minimal residual disease (MRD) (3,10,11). In most cases an elevated level of catecholamine production characterizes NB. Tyrosine hydroxylase (TH) catalyzes the first and rate-limiting step in the synthesis of catecholamines and therefore it can be used as a NB marker (2,3,12-14). Furthermore, a correlation between TH transcription and clinical outcome has been identified (2,12-14).
The Wilms tumor 1 gene, WT1, is located on chromosome 11p13 and encodes a zinc-finger protein that is associated in the development of some cases of Wilms tumor (15-17). WT1 is an archetypical dichotomous transcription factor in development and disease. WT1 plays an important role in regulating cellular growth and differentiation (17,18). Furthermore, it has been implicated as a diagnostic tool in monitoring MRD in leukemias (18-22). Although it is expressed at low levels in BM by normal progenitor cells, WT1 has been reported to be expressed at high levels in almost all hematological malignancies (18-20) and other solid tumors with their respective cell lines (15,18). Mutant or aberrantly expressed WT1 isoforms have also been described in desmoplastic SRCT, acute leukemias, mesothelioma, breast tumors and melanoma (18,23-25). During early development, WT1 is expressed in the brain and spinal cord, however its role in the malignancies that affect these tissues has not been previously investigated (18,25).
The objective of this study is to determine the potential association between WT1 and TH gene expression in high risk NB patients. To further elucidate the expression levels of both genes, samples were obtained at distinct clinical time points.
Materials and methods
Sample collection
After informed consent was given to use stored remains of DNA samples for research purposes, a total of 141 random samples from 34 eligible patients with the histological diagnosis of high risk NB by the Children’s Oncology Group (COG) criteria were collected. Including 27 samples at diagnosis [24 BM, 3 peripheral blood (PB)], 95 samples during treatment [62 BM, 3 PB, 30 peripheral blood stem cells (PBSC)], 13 post-transplant samples in remission (13 BM) and 6 samples at the time of clinical relapse (6 BM). Normal PB (n=19) was obtained from healthy volunteers and used as negative controls. Three NB cell lines (NGP, IMR-5 and LAN-5) were used for positive controls.
RNA preparation
Total RNA was extracted from BM, PB and PBSC samples using QIAampTM RNA Blood mini kit (QIAGEN Inc., Valencia, CA, USA) following the manufacturer’s procedure, for the determination of WT1 and TH mRNA gene expressions. The RNA concentration and purity was quantified by measuring the absorbance at 260 and 280 nm with a spectrophotometer (Pharmacia Biotech Gene Quant).
Quantitative RT-PCR
A two-step reverse transcriptase (RT)-PCR reaction was conducted in a 20 μL reaction volume containing 1 μg/μL of total RNA from each sample. Reaction buffer comprised of 4 μL of 25 nM MgCl2, 2 μL of 10x PCR buffer II, 8 μL dNTPs, 1 μL of Oligo d(T)16, 1 μL of RNase inhibitor, and 1 μL of Moloney Murine Leukemia Virus (MuLV) RT (Applied Biosystem, Foster City, CA, USA). Primer pairs were designed to locate specific nucleotide fragments for mRNA of both genes with the following sequences: TH (outer—sense) 5’-TGTCAGAGCTGGACAAGTGT-3’; (outer—antisense) 5’ GATATTGTCTTCCCGGTAGC-3’; (inner—sense) 5’ATCACCTGGTCACCAAGTTC-3’; (inner—antisense) 5’-GTGGTGTAGACCTCCTTCCA-3’and WT1 (outer—sense) 5’-GGCATCTGAGACCAGTGAGAA-3’; (outer—antisense) 5’ GAGAGTCAGACTTGAAAGCAGT-3’; (inner—sense) 5’-GCTGTCCCACTTACAGATGCA-3’; (inner—antisense) 5’-TCAAAGCGCCAGCTGGAGTTT-3’.
Reverse transcription was incubated in a 42 °C waterbath for 60 minutes, followed by denaturation at 96 °C in a thermal cycler for 10 minutes. 10 μL of the complementary DNA (cDNA) was amplified throughout the first round of PCR; it was then subjected for 30 cycles of initial denaturation for 5 minutes, amplification at 95 °C for 5 seconds, annealed at 55 °C for 5 seconds, and extension at 72 °C for 10 seconds. A second round of PCR was carried out by taking one μl of the amplified product from the first round and re-amplifying it with inner primers using the LightCycler SystemTM with SYBR-Green 1 RNA master mix reagent from Roche Diagnostics (Biochemica, Indianapolis, IN, USA). After PCR was completed, the concentration of target cells was calculated by the LightCycler software by extrapolation from the external standard curve. A standard curve was generated using a series of six ten-fold serial dilutions of NGP cell line, wherein sample results were deduced. The integrity of RNA was confirmed by amplification of the housekeeping gene GAPDH. RNA extracted from appropriate cell lines (NGP, IMR-5 and LAN-5) was used as a positive control for each fusion transcript. The identical procedure was performed for negative controls as well. Values of 1×100 ng/μL and above were considered positive for the expression of WT1 mRNA and TH mRNA, values between 9.9×10−1 to 6.5×10−1 ng/μL were considered weakly positive and any lower values were considered negative. The levels of WT1 and TH gene transcription in NGP were arbitrarily defined as 1.0. Specificity of PCR products was additionally verified in a 1.5% agarose gel electrophoresis. All RT-PCR experimentations were completed in triplicate and mean values were used.
Statistical analysis
Descriptive statistics were employed for the detection of association between TH and WT1 gene transcriptions as well as their individual significance in patient samples and controls, using Wilcoxon signed rank test analysis along with distinct column statistics were engaged at various clinical time points. For measurements of gene expression in patient samples in comparison to controls, the Wilcoxon rank sum test was used. Data analysis was done using GraphPad Prism version 5.0 software (GraphPad, La Jolla, CA, USA). P values of <0.05 was considered statistically significant.
Results
WT1 and TH gene expression in patients’ samples
As shown in Table 1 a total of 141 high risk NB patient samples were obtained and examined for TH and WT1 transcriptions at various clinical time points for the determination of association between the 2 genes. Paired levels of both gene transcriptions were extrapolated from a standard curve for the definitive measurements defined in Methods.
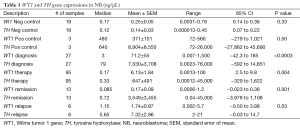
Full table
At diagnosis 93% (25/27) patient samples were detected to express positive and 7% (2/27) were considered weakly positive for TH gene expression. WT1 gene expression was also defined in the same manner with 70% (19/27) positive amplification and 15% (4/27) weakly positive. In addition 15% (4/27) of those patients’ samples were considered negative for WT1 transcript. When a comparison was made between TH and WT1 gene expressions at diagnosis (Figure 1A), a significant difference was found between the expression levels of the two genes (P<0.0003).
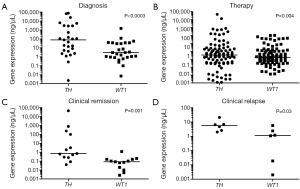
Throughout therapy at various time points high risk NB patient samples were acquired. When measuring TH gene expression 35% (33/95) in addition to 4% (4/95) of the samples expressed positive and weakly positive respectively, whereas 61% (58/95) expressed negative. Similarly, WT1 transcripts measured 70% (66/95) negative and 27% (26/95) positive with 3% (3/95) weakly positive. Significant difference (P=0.004) was measured amid WT1and TH transcriptions during therapy (Figure 1B).
During clinical remission values measuring TH and WT1 transcriptions varied. Overall TH gene expression was divided with 46% (6/13) negativity and 46% (6/13) positivity with 8% (1/13) weakly positive. In contrast, 92% (12/13) of WT1 expression was measured negative while only 8% (1/13) was positive. When evaluating the difference between the two varied gene expressions there was significance (P=0.001) established at clinical remission (Figure 1C).
On the other hand, patients in clinical relapse demonstrated 100% (6/6) positivity for TH transcription, in comparison to WT1 gene expression of 67% (4/6) positivity and 33% (2/6) negativity. Neither samples expressed weakly positive. Both gene expressions when compared at the time of clinical relapse (Figure 1D) illustrated significance (P=0.03).
Determination of WT1 and TH gene expression post HSCT
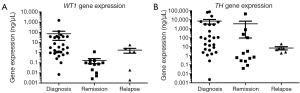
To evaluate the implication of both gene expressions in this high risk group, we measured each gene individually from patient samples at the time of initial diagnosis to post HSCT time points: clinical remission and clinical relapse. From diagnosis to the time of achieving clinical remission, patients samples produced a significant (P=0.0002) and non-significant difference (P=0.2) when measuring WT1 (Figure 2A) and TH (Figure 2B) respectively, while both genes were insignificant (P=0.06, P=0.09) when comparing samples from diagnosis to clinical relapse for transcriptions of their particular genes.

WT1 and TH gene expression in positive and negative controls
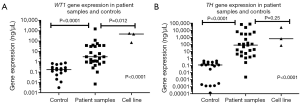
Overall, significance (P<0.0001) was determined when measuring the distinct level of gene expression of both WT1 (Figure 3A) and TH (Figure 3B) at the time of diagnosis in judgment to both negative and positive controls. Moreover, when comparing both gene expressions in negative and positive controls insignificance (P<0.33, P<0.50) was found correspondingly (Table 1).
Discussion
Dennis et al. (25) along with Wang et al. (18), found that WT1 gene is expressed in the brain as well as at detectable levels in human neural tumors, including brain tumors and NB. They conclude that WT1 gene does not play a significant role in the molecular etiology and pathogenesis of these tumors (18,25). In accordance to our study, using RT-PCR, we have detected varied levels of positive expression of WT1 in high-risk NB. Whereas others (23,24) have measured the same level of WT1 gene expression in NB at diagnosis using immunohistochemical staining. Irrespective of the technique that has been employed, WT1 positively expresses in NB.
As NB has a specific catecholamine metabolism, it may be useful to detect TH, which is the first rate-limiting enzyme of catecholamine synthesis (2,3,12-14). This inspired scientists to endeavor into measuring the level of TH gene expression in NB. Indeed, increased expression of the TH gene is indicative of the presence of NB cells, as seen in several studies (26-29). In addition Ootsuka et al. (3) have found that it is a specific marker for NB in comparison to other SRCTs. Thus, TH has been used for the detection of MRD in NB (30-36). Accordingly, in our study, for the assessment of WT1 gene expression, TH was employed as a known marker of NB.
Overall, although there was a significant difference found between both gene transcriptions throughout the study, WT1 demonstrated similar gene expression to TH by the insignificance measured in both positive and negative controls (Table 1). At the time of diagnosis and during therapy patient samples expressed an equal WT1/TH ratio of 0.01, while samples post-transplant in remission expressed a decreased ratio of 0.00005. WT1 gene was negative and significantly lower than TH, which remained highly positive, in patient samples measured from BM in clinical remission. Based on our previous studies (20-22) this exemplifies that WT1 is a marker of proliferation and that these BM samples from patients post-transplant illustrate a non-proliferative metastatic state in high risk NB. Additionally, spontaneous or chemotherapy-induced differentiation of a malignant neuroblastic tumor, including NB, is known to mature into ganglioneuroma (GN) (6-9), which is the most differentiated neuroblastic tumors. This non-proliferative state of GN can be seen histologically in this setting. Consequently, it is possible that WT1 gene expression may be negative. This may explain why even in clinical presence of tumor cells the WT1 gene may not be expressed post-transplant. When comparing samples from diagnosis to remission, WT1 and TH transcription illustrated significant and insignificant difference respectively. The insignificance found in TH transcription contributes to TH being positively expressed in normal and malignant neuronal tissue (37,38).
WT1 can be compared to TH and that at the time of clinical remission both gene expression vary due to their independent physiology in NB. Moreover, these findings suggest that WT1 is a surrogate marker of proliferation in this high-risk group. The use of TH as an MRD marker in NB is hampered by its illegitimate expression in normal hematological cells (4,28,36-40). Thus in order to avoid false negative results, WT1 should be adjuvant to TH for the detection of MRD in this high-risk group. The need to conduct larger prospective studies is pivotal for the proper understanding of what role does the (WT1) plays in NB.
Acknowledgements
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Cheung NK, Dyer MA. Neuroblastoma: developmental biology, cancer genomics and immunotherapy. Nat Rev Cancer 2013;13:397-411. [PubMed]
- Avigad S, Feinberg-Gorenshtein G, Luria D, et al. Minimal residual disease in peripheral blood stem cell harvests from high-risk neuroblastoma patients. J Pediatr Hematol Oncol 2009;31:22-6. [PubMed]
- Ootsuka S, Asami S, Sasaki T, et al. Useful markers for detecting minimal residual disease in cases of neuroblastoma. Biol Pharm Bull 2008;31:1071-4. [PubMed]
- Stutterheim J, Gerritsen A, Zappeij-Kannegieter L, et al. Detecting minimal residual disease in neuroblastoma: the superiority of a panel of real-time quantitative PCR markers. Clin Chem 2009;55:1316-26. [PubMed]
- Stutterheim J, Zappeij-Kannegieter L, Ora I, et al. Stability of PCR targets for monitoring minimal residual disease in neuroblastoma. J Mol Diagn 2012;14:168-75. [PubMed]
- Adam O, Boia ES, Ilie R, et al. Ganglioneuroma. Jurnalul Pediatrului 2007;10:48-51.
- Iwanaka T, Yamamoto K, Ogawa Y, et al. Maturation of mass-screened localized adrenal neuroblastoma. J Pediatr Surg 2001;36:1633-6. [PubMed]
- Przkora R, Perez-Canto A, Ertel W, et al. Ganglioneuroma: primary tumor or maturation of a suspected neuroblastoma? Eur Spine J 2006;15:363-5. [PubMed]
- Rozmus J, Langer M, Murphy JJ, et al. Multiple persistent ganglioneuromas likely arising from the spontaneous maturation of metastatic neuroblastoma. J Pediatr Hematol Oncol 2012;34:151-3. [PubMed]
- Tchirkov A, Paillard C, Halle P, et al. Significance of molecular quantification of minimal residual disease in metastatic neuroblastoma. J Hematother Stem Cell Res 2003;12:435-42. [PubMed]
- Reynolds CP. Detection and treatment of minimal residual disease in high-risk neuroblastoma. Pediatr Transplant 2004;8 Suppl 5:56-66. [PubMed]
- Parareda A, Gallego S, Roma J, et al. Prognostic impact of the detection of microcirculating tumor cells by a real-time RT-PCR assay of tyrosine hydroxylase in patients with advanced neuroblastoma. Oncol Rep 2005;14:1021-7. [PubMed]
- Naito H, Kuzumaki N, Uchino J, et al. Detection of tyrosine hydroxylase mRNA and minimal neuroblastoma cells by the reverse transcription-polymerase chain reaction. Eur J Cancer 1991;27:762-5. [PubMed]
- Lee ST, Suh YL, Ko YH, et al. Measurement of tyrosine hydroxylase transcripts in bone marrow using biopsied tissue instead of aspirates for neuroblastoma. Pediatr Blood Cancer 2010;55:273-8. [PubMed]
- Sugiyama H. Wilms’ tumor gene WT1: its oncogenic function and clinical application. Int J Hematol 2001;73:177-87. [PubMed]
- Sugiyama H. WT1 (Wilms' tumor gene 1): biology and cancer immunotherapy. Jpn J Clin Oncol 2010;40:377-87. [PubMed]
- Scharnhorst V, van der Eb AJ, Jochemsen AG. WT1 proteins: functions in growth and differentiation. Gene 2001;273:141-61. [PubMed]
- Wang J, Oue T, Uehara S, et al. The role of WT1 gene in neuroblastoma. J Pediatr Surg 2011;46:326-31. [PubMed]
- Ogawa H, Tamaki H, Ikegame K, et al. The usefulness of monitoring WT1 gene transcripts for the prediction and management of relapse following allogeneic stem cell transplantation in acute type leukemia. Blood 2003;101:1698-704. [PubMed]
- Kletzel M, Olzewski M, Huang W, et al. Utility of WT1 as a reliable tool for the detection of minimal residual disease in children with leukemia. Pediatr Dev Pathol 2002;5:269-75. [PubMed]
- Olszewski M, Huang W, Chou PM, et al. Wilms' tumor 1 (WT1) gene in hematopoiesis: a surrogate marker of cell proliferation as a possible mechanism of action? Cytotherapy 2005;7:57-61. [PubMed]
- Olszewski M, Chou PM, Huang W, et al. Correlation of minimal residual disease by assessing Wilms tumor gene expression and engraftment by variable number of tandem repeats in children with leukemia posthematopoietic stem cell transplantation. Pediatr Dev Pathol 2006;9:203-9. [PubMed]
- Sebire NJ, Gibson S, Rampling D, et al. Immunohistochemical findings in embryonal small round cell tumors with molecular diagnostic confirmation. Appl Immunohistochem Mol Morphol 2005;13:1-5. [PubMed]
- Barnoud R, Sabourin JC, Pasquier D, et al. Immunohistochemical expression of WT1 by desmoplastic small round cell tumor: a comparative study with other small round cell tumors. Am J Surg Pathol 2000;24:830-6. [PubMed]
- Dennis SL, Manji SS, Carrington DP, et al. Expression and mutation analysis of the Wilms' tumor 1 gene in human neural tumors. Int J Cancer 2002;97:713-5. [PubMed]
- Fukuda M, Miyajima Y, Miyashita Y, et al. Disease outcome may be predicted by molecular detection of minimal residual disease in bone marrow in advanced neuroblastoma: a pilot study. J Pediatr Hematol Oncol 2001;23:10-3. [PubMed]
- Horibe K, Fukuda M, Miyajima Y, et al. Outcome prediction by molecular detection of minimal residual disease in bone marrow for advanced neuroblastoma. Med Pediatr Oncol 2001;36:203-4. [PubMed]
- Lambooy LH, Gidding CE, van den Heuvel LP, et al. Real-time analysis of tyrosine hydroxylase gene expression: a sensitive and semiquantitative marker for minimal residual disease detection of neuroblastoma. Clin Cancer Res 2003;9:812-9. [PubMed]
- Corrias MV, Haupt R, Carlini B, et al. Peripheral blood stem cell tumor cell contamination and survival of neuroblastoma patients. Clin Cancer Res 2006;12:5680-5. [PubMed]
- Burchill SA, Bradbury FM, Smith B, et al. Neuroblastoma cell detection by reverse transcriptase-polymerase chain reaction (RT-PCR) for tyrosine hydroxylase mRNA. Int J Cancer 1994;57:671-5. [PubMed]
- Corrias MV, Faulkner LB, Pistorio A, et al. Detection of neuroblastoma cells in bone marrow and peripheral blood by different techniques: accuracy and relationship with clinical features of patients. Clin Cancer Res 2004;10:7978-85. [PubMed]
- Miyajima Y, Kato K, Numata S, et al. Detection of neuroblastoma cells in bone marrow and peripheral blood at diagnosis by the reverse transcriptase-polymerase chain reaction for tyrosine hydroxylase mRNA. Cancer 1995;75:2757-61. [PubMed]
- Oltra S, Martinez F, Orellana C, et al. The doublecortin gene, a new molecular marker to detect minimal residual disease in neuroblastoma. Diagn Mol Pathol 2005;14:53-7. [PubMed]
- Swerts K, De Moerloose B, Dhooge C, et al. Potential application of ELAVL4 real-time quantitative reverse transcription-PCR for detection of disseminated neuroblastoma cells. Clin Chem 2006;52:438-45. [PubMed]
- Tchirkov A, Paillard C, Halle P, et al. Significance of molecular quantification of minimal residual disease in metastatic neuroblastoma. J Hematother Stem Cell Res 2003;12:435-42. [PubMed]
- Träger C, Kogner P, Lindskog M, et al. Quantitative analysis of tyrosine hydroxylase mRNA for sensitive detection of neuroblastoma cells in blood and bone marrow. Clin Chem 2003;49:104-12. [PubMed]
- Weihe E, Depboylu C, Schütz B, et al. Three types of tyrosine hydroxylase-positive CNS neurons distinguished by dopa decarboxylase and VMAT2 co-expression. Cell Mol Neurobiol 2006;26:659-78. [PubMed]
- Young HM, Ciampoli D, Hsuan J, et al. Expression of Ret-, p75(NTR)-, Phox2a-, Phox2b-, and tyrosine hydroxylase-immunoreactivity by undifferentiated neural crest-derived cells and different classes of enteric neurons in the embryonic mouse gut. Dev Dyn 1999;216:137-52. [PubMed]
- Viprey VF, Corrias MV, Kagedal B, et al. Standardisation of operating procedures for the detection of minimal disease by QRT-PCR in children with neuroblastoma: quality assurance on behalf of SIOPEN-R-NET. Eur J Cancer 2007;43:341-50. [PubMed]
- Kuçi Z, Seitz G, Kuçi S, et al. Pitfalls in detection of contaminating neuroblastoma cells by tyrosine hydroxylase RT-PCR due to catecholamine-producing hematopoietic cells. Anticancer Res 2006;26:2075-80. [PubMed]


