Viral etiology and atopic characteristics in high-risk asthmatic children hospitalized for lower respiratory tract infection
Introduction
The global prevalence of asthma in children is increasing year by year, bringing a heavy economic burden to families and society (1). Previous studies at home and abroad classified asthma into early transient wheezing, early-onset persistent wheezing (onset before 3 years old) and late-onset wheezing/asthma (2,3). A useful means for confirming the diagnosis of asthma in children aged ≤5 years old is a trial of treatment with short-acting bronchodilators and inhaled corticosteroids. Marked clinical improvement during the treatment, and deterioration when treatment is stopped, supports a diagnosis of asthma. The interaction between atopy and viral infections is complex. The atopic state influences lower airway responses to viral infections and viral infections can influence the development of allergic sensitization, especially when individuals are exposed simultaneously to both (2). Treatment options for childhood asthma can be classified into standard therapies, targeted therapies and other therapies. Most children with asthma are well-controlled by standard therapies recommended by GINA. For children treated with high-dose inhaled corticosteroids (ICS), the addition of long-acting beta agonists and leukotriene receptors can be considered, it can also be treated with oral prednisolone; however, a small number of children with refractory asthma require biological targeted therapy, which can reduce acute asthma attacks and hormone dosages, and improve symptoms and lung function (4).
Therefore, early recognition and high-risk factors for wheezing contributes an important role on the prevention of asthma. About 40–50% of infants have had at least one episode of wheezing, and 40–50% of children have later wheezing through at least 6 years of age, some of whom later develop asthma. Respiratory viral infection and allergic reactions (atopic constitution) are common causes of wheezing and repeated wheezing in infants and young children (5).
Respiratory viruses play a vital role in the development and exacerbation of obstructive respiratory diseases in children. Rossi et al. (6) found that the susceptibility to HRV-induced bronchiolitis and subsequent wheezing is linked to individual predisposition since it is often associated with a family or personal history of asthma/atopy. However, little research on this link has been conducted in China, so the present work studied infants and young children in China. We performed real-time fluorescent polymerase chain reaction (PCR) and ImmunoCAP assays to determine the distribution of viral pathogens and allergen sensitivities in high-risk asthmatic children.
An assessment of inflammation is becoming standard practice in the clinical work-up of children with persistent asthma. Konradsen et al. (7) found that assessment of both local and systemic Th2-mediated inflammation via the analysis of readily available biomarkers, including a fractional exhaled nitric oxide (FeNO) and blood eosinophils, had a high predictive value for the identification of children with the highest asthma morbidity. A population-based study recently found that the prevalence of wheeze, asthma, and exacerbations increased independently and additively with increasing levels of FeNO and blood eosinophils (8).
Viral pathogens, allergens, and levels of FeNO are good predictors for high-risk asthmatic children developing to asthma or not, and the connections among them are not defined. This study was conducted to investigate these unanswered questions, providing help for early diagnosis and prevention of high-risk asthmatic children.
We present the following article in accordance with the MDAR reporting checklist (available at http://dx.doi.org/10.21037/tp-20-165).
Methods
Patients
Pediatric patients, hospitalized for lower respiratory tract infection (LRTI) with wheezing, were included in a prospective study conducted from April 2016 until August 2017 in the Department of Respiratory Medicine, Children’s Hospital of Soochow University. The group of these infants and young children was called “high-risk of asthma infants.” The inclusion criteria were (I) aged younger than 3 years, (II) having a first or second episode of wheezing, and (III) meeting at least one of the following conditions: (I) family history of at least one first-degree relative with asthma; (II) atopy (i.e., allergic rhinitis, eczema, or food allergies diagnosed by a medical doctor); (III) elevated peripheral blood eosinophil level (>4%); or (IV) inhalant or food allergen sensitization of level II or above in a previous allergen test. The exclusion criteria were (I) having various congenital diseases or birth defects, (II) airway malformations or pulmonary dysplasia, (III)pulmonary interstitial disease or severe pneumonia, (IV)other non-respiratory diseases causing wheezing, and (V) suffering from cardiovascular disease, liver, kidney or metabolic disease.
Similarly, from April 2016 to August 2017, non-asthmatic and non-allergic healthy children with normal cardiac ultrasound carried out in the Echocardiography clinic were recruited as the control group for the measurement of FeNO.
The experimental protocols were approved by the ethics committee of the Children’s Hospital of Soochow University (NO.: 2015LW006). Written informed consent of the guardians of included infants was obtained before enrollment. All procedures performed in this study involving human participants were in accordance with the Declaration of Helsinki (as revised in 2013).
Sample collection
Nasopharyngeal aspirates were collected from all patients during the first 24 h following hospital admission. Laboratory tests were performed on the same day in which the specimens were obtained. Sample qualification criteria included a white blood cell count of >25 cells/low power field and a squamous epithelial cell count of <10 cells/low power field. Samples were stored at 2–8 °C until analysis.
Extraction of nucleic acids
Rapid purification of high-quality viral nucleic acid (RNA/DNA) was performed using a QIAamp MinElute Virus Spin Kit (Qiagen, Hilden, Germany) (9).
Viral nucleic acid was extracted from 200 µL of nasopharyngeal aspirate specimens. The procedure was composed of four steps (lyse, bind, wash, elute). Samples were lysed under highly denaturing conditions at elevated temperatures. The lysis was performed in Qiagen Protease and Buffer AL, which together ensure the inactivation of RNases. Binding conditions were adjusted by adding ethanol to allow for the optimal binding of viral RNA and DNA to the membrane. Nucleic acids remained bound to the membrane, while contaminants were efficiently washed away during three wash steps. Elution was performed using Buffer AVE.
The QIAamp MinElute Virus Spin process was fully automated on the QIAcube (QIAGEN, Shanghai, China) following the manufacturer’s instructions.
Virus detection
The respiratory pathogens kit from Da An Gene was used for virus detection. This method can detect and characterize the seven most common types of human viruses that cause respiratory infection: adenovirus (ADV); human bocavirus (HBoV); influenza type A (Flu-A); parainfluenza virus (PIV); human rhinovirus (HRV); respiratory syncytial virus (RSV), and human metapneumovirus (hMPV).
This kit, with real-time fluorescent PCR, uses a conserved region of the viral genome as a target region. It designs specific primers and fluorescent probes to quantitatively detect viral DNA/RNA via in vitro PCR amplification.
Reaction conditions: ADV: 50 °C 15 min, 95 °C 15 min, 95 °C 15 s, 55 °C 45 s, 40 cycles; HBoV: 50 °C 2 min, 95 °C 15 min; 95 °C 15 s, 55 °C 45 s, 40 cycles; Flu-A: 50 °C 15 min, 95 °C 15 min, 95 °C 15 s, 58 °C 45 s, 45 cycles; PIV: 50 °C 15 min, 95 °C 15 min, 95 °C 15 s, 55 °C 45 s, 45 cycles; HRV: 50 °C 15 min, 95 °C 15 min, 94 °C 15 s, 55 °C 45 s, 40 cycles; RSV: 40 °C 30 min, 94 °C 3 min, 93°C 15 s, 55 °C 45 s, 10 cycles; 93 °C 15 s, 55 °C 45 s, 30 cycles; hMPV: 50 °C 15 min, 95 °C 15 min, 95 °C 15 s, 55 °C 45 s, 40 cycles. According to the fluorescence curve, the results are displayed by Ct value, and the computer automatically analyzes the results.
Allergic sensitization
The total serum concentration of immunoglobulin E (T-IgE, kU/L) and the levels of IgE antibodies directed against the PhadiatopÒa mixture of common inhaled allergens (dust mites, pollen, fungi, and animal dander) and against food allergens (fx5Ò = chicken protein and milk) were determined using the ImmunoCAPÒ System (Phadia AB, Uppsala, Sweden). Atopy was defined as a level of IgE antibodies toward PhadiatopÒ or fx5Ò of ≥0.35 kU/L.
The following allergens were included in the test: dust mites (Dermatophagoides farinae and Dermatophagoides pteronyssinus), pollen (spring and summer pollen, summer and autumn pollen), fungi (Aspergillus fumigatus and Alternaria), animal dander (cat and dog), chicken protein, and milk. The laboratory analyses were performed in a blinded manner, and the results were sent back to the investigating physician.
FeNO
A NIOXÒ analyzer (Aerocrine AB, Solna, Sweden) was used to measure the fraction of FeNO (10). FeNO is presented as absolute values in parts per billion (p.p.b.) or as height-adjusted FeNO values (FeNO%) (11). All participants were calmed down well before the examination, and they did not have any strenuous exercise or intake of nitrogen-rich foods (including sausages, animal offal, lettuce, and spinach) or caffeinated drinks within the 2 h before the test.
Statistical analysis
Data were stored in a Microsoft Excel-supported database, and statistical analyses were performed using SPSS statistics version 20.0. The measurement data with a non-normal distribution are expressed here as the median (interquartile spacing) [M (P25, P75)]. A rank-sum test was used to analyze between-group differences in T-IgE among groups on subject age, virus detection, and wheezing frequency and to analyze between-group differences in FeNO among groups on wheezing frequency. Qualitative data are presented here as the frequency distribution of percentages in each category. A chi-square test was used to evaluate associations between inhaled allergens and food allergens among groups on age, virus detection, and wheezing frequency. Results with a P value of <0.05 were considered significant.
Results
Demographic data
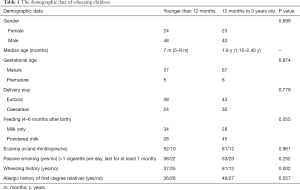
The current study included 135 children with an elevated risk of asthma. Among them, 78 (57.78%) patients were male, and 57 (42.22%) were female (male-to-female ratio: 1.37:1); the median age was 12 months (interquartile range, 7–21 months). Of the 62 (45.93%) children aged less than 12 months, the median age was 7 months (interquartile range, 5–9 months). The median age of the 73 (54.07%) children aged 1–3 years old was 1.9 years (interquartile range, 1.15–2.40 years). Demographic data were collected uniformly from all included children (Table 1).
Full table
Detection and distribution of viruses
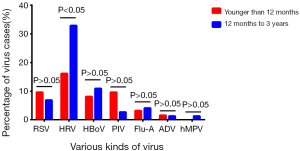
Nasopharyngeal aspirate samples from 135 pediatric patients with an elevated risk of asthma were analyzed using PCR. Viral nucleic acids were positively detected in 67 (49.63%) samples. The most frequently detected viruses were HRV (34 cases), HBoV (13 cases), RSV (11 cases), Flu-A (8 cases), PIV (5 cases), ADV (2 cases), and hMPV (1 case). Seven cases had coinfections of two viruses.
There were 62 (45.93%) subjects aged younger than 12 months and 73 (54.07%) subjects aged 1–3 years old. Among infants aged younger than 12 months, the most common viruses were HRV (10 cases), RSV (6 cases), and PIV (6 cases); among those aged 1–3 years old, the most common viruses were HRV (24 cases), HBoV (8 cases), and RSV (5 cases). The older subset of study subjects (aged 1–3 years) was more susceptible to HRV infection compared with the younger subset (aged <12 months) (P=0.03; Figure 1).
Detection and distribution of allergens
Allergic sensitization was detected in 59.26% of patients; 88.75% and 43.75% of patients had positive reactions to food and inhalant allergens, respectively. Among the tested inhalant allergens, sensitivity to dust mites, fungi, pollen, and animal dander was found in 77.14%, 37.14%, 25.71%, and 8.57% of patients, respectively. Regarding allergic sensitivity to multiple inhalant allergens, 8.57% and 14.29% of patients had positive reactions to two and three allergens, respectively; no patients reacted to four allergens. Among the tested food allergens, sensitivity to chicken protein, milk, and both allergens was detected in 73.24%, 67.61%, and 42.25% of subjects, respectively. Additionally, 32.50% of patients showed positive reactions against both food and inhalant allergens, and the rate of positive reaction against food allergens was significantly higher than that against inhalant allergens.
Compared with children aged 1–3 years, infants aged younger than 12 months had a significantly higher positive reaction rate against inhalant allergens (P<0.05) and a lower level of T-IgE (P<0.05), whereas there were no significant differences in reactivity to food allergens between these two groups (Table 2).

Full table
Relationship between viral detection and allergen sensitivity
Regarding the groups of patients with positive detection of one or more of the seven common respiratory viruses, no significant differences were observed among the positive rates of reactivity against food or inhalant allergens or the levels of T-IgE. However, the larger group of patients in which any virus was detected (virus-detected group) had a significantly higher positive rate of sensitivity against inhalant allergens compared with the group of patients in which no viruses were detected (virus-undetected group) (P=0.012). However, there were no significant differences in the food allergen sensitivity rate or the T-IgE level between these two groups (Table 3).

Full table
FeNO
A group of control subjects was recruited for comparison to investigate FeNO in the high-risk asthmatic children (observation group). The control group consisted of 200 non-asthmatic, non-allergic healthy children with a male (105 cases)-to-female (95 cases) ratio of 1.11:1 and a median age of 12 months (6.25–24.00 months). The sex ratio and average age were comparable between the observation group and the control group (P>0.05). The FeNO values in the observation group [3.80 (2.90, 8.33) µg/L] were significantly lower than those in the control group [9.60 (5.60, 16.90) µg/L] (P=0.001). The FeNO values in children with RSV, HRV, PIV, Flu-A, ADV, hMPV, and viral coinfection were 3.55 (1.78, 8.25) µg/L, 3.80 (3.15, 7.60) µg/L, 5.70 (1.88, 11.33) µg/L, 4.80 (3.50, 6.90) µg/L, 3.60 (5.12) µg/L, 3.50 (4.87) µg/L, and 3.30 (2.40, 3.40) µg/L, respectively, all of which were significantly lower than those in the control group children. However, the mean FeNO value of children with HBoV infection was 13.40 (7.08, 22.98) µg/L, significantly higher than that in the control group children.
Comparison of allergens and FeNO among children with different wheezing frequencies
The 135 high-risk asthmatic children were split into two groups on their wheezing frequency (first-wheezing group and second wheezing group). Compared with the first-wheezing group, the second-wheezing group had a higher rate of positive reaction against dust mites and fungi and a higher T-IgE level (P=0.045) (Table 4); however, the rates of positive reaction to animal dander, pollen, chicken protein, and milk and the levels of T-IgE and FeNO were not significantly different between the two groups (P>0.5).
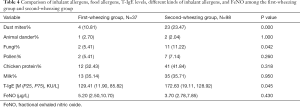

Full table
Discussion
In recent years, the morbidity of asthmatic diseases and later wheezing has been increasing in young children. Because there are many various causes of wheezing and the detection technology used for diagnosing asthma in older children or adults is unsuitable for use in young children, the diagnosis of infantile asthma is a massive problem for clinicians and is an area of active research. It is an arduous task to set up and evaluate a series of auxiliary detection techniques and standards, including infantile tidal breathing lung function, impulse oscillometry lung function, FeNO, and calculations of sputum eosinophils. However, it is of practical significance to study the clinical characteristics, risk factors (including allergies), and pathogenic characteristics of an asthma attack in infants and young children, to devise methods for the early prevention and treatment of asthma. Many previous clinical observations have revealed that viral infection is the most common cause of infantile and young children’s wheezing (12).
Our research here used real-time fluorescence PCR to detect viral nucleic acid in nasopharyngeal aspirates from children at elevated risk of developing asthma. The total positive rate of virus detection in our subjects was 49.63%, which is almost consistent with the values of 51.00–59.7% (13,14) reported in the medical literature from outside China. HRV was the most frequently detected virus in our samples; this virus is the leading cause of respiratory diseases (15-18).
In a birth cohort study of children at high risk of developing asthma in Australia (19), significant correlations were found between wheezing during infancy, HRV infection within the first year of life, and the incidence of asthma at the age of 5 years. Meanwhile, Kotaniemi-Syrjänen et al. (20) also reported that HRV infection was more prevalent than any other viral infection in children aged under 2 years who were hospitalized for wheezing; children infected with HRV had a risk of developing asthma four times higher than that in those not infected with HRV. Due to the imbalance of Th1/Th2 in atopic children, they are more susceptible to HRV infection. Th1 dominance involves a down-regulated expression of cytokines associated with viral replication, including IL-12, IFN-α, IFN-γ, and IFN-λ. Among asthmatic patients, the expression of these factors is lower, shifting the balance toward Th2 immunity, which is beneficial for the replication of HRV. Therefore, early HRV infection is a high-risk factor for asthma (21). Viral coinfection is also a risk factor for infantile episodes of wheezing, but the details of this influence are still unclear (22). An infantile viral infection is closely related to wheezing (23). Rossi et al. (6) proposed that early RSV infection is a risk factor for subsequent wheezing and even asthma in childhood. RSV likely serves as an “inducer” instead of a “trigger” in most people; however, it seems to serve as a “trigger” rather an “inducer” in predisposed individuals.
The measurement of Th2-mediated biomarkers, including FeNO, has become a common practice in the clinical work-up of children with persistent asthma (7,24). Many studies have shown that FeNO can be used as a marker reflecting eosinophilic respiratory inflammation (25), and FeNO measurement has been widely recommended by many guidelines, both in China and abroad, to assist in the diagnosis of asthma, assessment of the patient’s condition, and prediction of patient response to hormone therapy. Unexpectedly, the present study found that FeNO was significantly lower in infants and young children at elevated risk of developing asthma than in healthy children. Some scholars have pointed out that the acute phase of RSV infection does not include eosinophilic respiratory inflammation (26); additionally, virus-induced wheezing is most closely related to neutrophil respiratory inflammation (27), which leads to a temporary decline in FeNO. The group of children at high risk of developing asthma in this study who were found to be infected with a common respiratory virus (i.e., RSV, HRV, Flu-A, PIV, ADV, or hMPV) or coinfected with multiple viruses may have been experiencing virus-induced transient wheezing and are not asthmatic because wheezing is not a manifestation of eosinophilic respiratory tract inflammation. Notably, Konradsen et al. (7) found that when the interference of allergic factors was excluded, the FeNO of children with viral LRTI, increased after the viral infection was controlled, regardless of whether it was accompanied by wheezing or not. This is like the “rebound phenomenon” observed by Gadish et al. (26). In the present study, the observed low FeNO level after viral infection may be due to the restriction imposed by bronchial spasms in the acute stage of wheezing. Because our study did not include follow-up visits with the subjects, we were unable to investigate the “rebound phenomenon” described above; future work could reassess this group of subjects to investigate this issue.
Global Strategy for Asthma Management and Prevention updated 2018 (GINA2018) found that for preschool children with an LRTI exhibiting recurrent cough and wheezing, the FeNO continued to rise for more than four weeks, and this rise could be used to predict the occurrence of asthma (2). Therefore, continuous dynamic monitoring of FeNO changes has some value for the early prediction of childhood asthma. Here, the FeNO level was higher in subjects infected with HBoV than in healthy controls. Huang et al. (28) found that HBoV infection caused pathological effects on bronchial epithelial cells, leading to an increase in the production of inducible nitric oxide synthase (iNOS) and FeNO; however, the pathogenic mechanism needs to be further studied.
Having an atopic constitution is an independent risk factor for the development of persistent asthma (29). Phadiatop + Fx5 combined detection can help effectively identify IgE-mediated diseases in infants and children and is an ideal method for in vitro allergen screening. Here, 59.26% of subjects at elevated risk of developing asthma were identified as having allergic sensitization, indicating that this group of asthmatic children had a high proportion of atopic constitutions. Additionally, we found that the positive rates of sensitivity to inhalant allergens and food allergens in infants with second wheezing were significantly higher than those in children with first wheezing, suggesting that repeated wheezing is related to having an atopic constitution.
In this study, the rate of sensitivity to food allergens was significantly higher than that of inhaled allergens, which may be due to the reduced function of the intestinal tract barrier in infants and young children. Food allergens tend to infiltrate into the blood circulation through the loosely connected epithelial cell gap, thus causing allergy symptoms of the skin and gastrointestinal and respiratory tracts. Many studies have shown that the most common inhaled allergens in children with bronchial asthma are dust mites and fungi. We found that the rates of sensitivity to inhaled allergens from high to low were 33.75% for dust mites, 16.25% for fungi, 11.25% for pollen, and 3.75% for animal dander. The rates of sensitivity to dust mites and fungi in infants with second wheezing were higher than those in children with first wheezing. Our data indicate that the positive rate of sensitivity to inhaled allergens in the group of infants aged less than 12 months was higher than that in the group of children aged 1–3 years. The reason for this is difficult to determine, but it could be related to the selected subjects of this study being “high-risk” children. Previous work has shown that early exposure to inhaled allergens and repeated wheezing during childhood are high-risk factors for developing asthma or other atopic disorders (30); these are also the significant risk factors in asthma prediction index (API). Also, due to the immaturity of the immune system in infants and young children and the different load intensity of allergen exposure in the environment, there may be some differential changes over time, so assays against Phadiatop + Fx5 should be repeated 3 to 6 months later if necessary (31). The T-IgE level in the group aged 1–3 years old was higher than that in the group aged <12 months old. As the immune system develops with age, increased exposure to allergens in the environment stimulates the body to produce more IgE. The positive rate of sensitivity to inhaled allergens in the virus-detected group was higher than that in the virus-undetected group. Therefore, the relationship between viral infection and sensitivity to inhaled allergens should be further studied to explore the associated mechanism.
Conclusions
In conclusion, early HRV infection and sensitivity to inhalant allergens are high-risk factors for asthma, and having an atopic constitution is closely related to respiratory viral infection in infants and young children. Therefore, it is of practical significance to understand the risk factors of infants and young children’s wheezing (including pathogenic characteristics and atopic constitutions), as this could aid in identifying asthma and beginning appropriate interventions at an early stage. The value of a single FeNO measurement is limited, so continuous dynamic monitoring may be more meaningful in predicting the occurrence of asthma; however, FeNO measurement may be a useful technique for evaluating the treatment and prognosis of asthma.
Acknowledgments
We thank Katie Oakley, PhD, from LiwenBianji, Edanz Editing China (www.liwenbianji.cn/ac), for editing the English text of a draft of this manuscript.
Funding: This work was supported by the National Natural Science Foundation of China [81771676, 81701948, 81870006, 81970027, 81971490]; Science and Technology Project of Suzhou [SS201765, SS201869]; Social Development Projects of Jiangsu Province [BE2016676, BE2019671]; and Key Lab of Respiratory Disease of Suzhou [SZS201714].
Footnote
Reporting Checklist: The authors have completed the MDAR reporting checklist. Available at http://dx.doi.org/10.21037/tp-20-165
Data Sharing Statement: Available at http://dx.doi.org/10.21037/tp-20-165
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/tp-20-165). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All procedures performed in this study involving human participants were in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the ethics committee of the Children’s Hospital of Soochow University (NO.: 2015LW006) and informed consent was taken from all the patients.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Ebmeier S, Thayabaran D, Braithwaite I, et al. Trends in international asthma mortality: analysis of data from the WHO Mortality Database from 46 countries (1993-2012). Lancet 2017;390:935-45. [Crossref] [PubMed]
- Lin J, Xing B, Chen P, et al. Chinese expert consensus-based guideline on assessment and management of asthma exacerbation. J Thorac Dis 2019;11:4918-35. [Crossref] [PubMed]
- Chen X, Qiu C. Respiratory tract mucous membrane microecology and asthma. Ann Transl Med 2019;7:495. [Crossref] [PubMed]
- Weir E, Paton J. Mepolizumab in adolescents with severe eosinophilic asthma not eligible for omalizumab: one center's early clinical experience. J Asthma 2020;57:521-4. [Crossref] [PubMed]
- Weiss LN. The diagnosis of wheezing in children. Am Fam Physician 2008;77:1109-14. [PubMed]
- Rossi GA, Colin AA. Infantile respiratory syncytial virus and human rhinovirus infections: respective role in inception and persistence of wheezing. Eur Respir J 2015;45:774-89. [Crossref] [PubMed]
- Konradsen JR, Skantz E, Nordlund B, et al. Predicting asthma morbidity in children using proposed markers of Th2-type inflammation. Pediatr Allergy Immunol 2015;26:772-9. [Crossref] [PubMed]
- Andrei M, Fonseca JOA, Tiago J, et al. Exhaled nitric oxide levels and blood eosinophil counts independently associate with wheeze and asthma events in National Health and Nutrition Examination Survey subjects. J Allergy Clin Immunol 2013;132:821-7.e1-5.
- Yan XL, Li YN, Tang YJ, et al. Clinical characteristics and viral load of respiratory syncytial virus and human metapneumovirus in children hospitaled for acute lower respiratory tract infection. J Med Virol 2017;89:589-97. [Crossref] [PubMed]
- Peltier RE. ATS/ERS recommendations for standardized procedures for the online and offline measurement of exhaled lower respiratory nitric oxide and nasal nitric oxide, 2005. Am J Respir Crit Care Med 2005;171:912. [Crossref] [PubMed]
- Malmberg LP, Pet Ys T, Haahtela T, et al. Exhaled nitric oxide in healthy nonatopic school-age children: determinants and height-adjusted reference values. Pediatr Pulmonol 2006;41:635-42. [Crossref] [PubMed]
- Connors TJ, Baird JS, Yopes MC, et al. Developmental Regulation of Effector and Resident Memory T Cell Generation during Pediatric Viral Respiratory Tract Infection. J Immunol 2018;201:432-9. [Crossref] [PubMed]
- Leung TF, Man YT, Yeung ACM, et al. Multiplex Molecular Detection of Respiratory Pathogens in Children With Asthma Exacerbation. Chest 2010;137:348-54. [Crossref] [PubMed]
- Moreno-Valencia Y, Hernandez-Hernandez VA, Romero-Espinoza JAI, et al. Detection and characterization of respiratory viruses causing acute respiratory illness and asthma exacerbation in children during three different seasons (2011-2014) in Mexico City. Influenza Other Respir Viruses 2015;9:287-92. [Crossref] [PubMed]
- Tsatsral S, Xiang Z, Fuji N, et al. Molecular Epidemiology of the Human Rhinovirus Infection in Mongolia during 2008-2013. Jpn J Infect Dis 2015;68:280-7. [Crossref] [PubMed]
- Lu QB, Wo Y, Wang LY, et al. Molecular epidemiology of human rhinovirus in children with acute respiratory diseases in Chongqing, China. Sci Rep 2014;4:6686. [Crossref] [PubMed]
- Kainulainen V, Elf S, Susi P, et al. Detection of human rhinoviruses by reverse transcription strand invasion based amplification method (RT-SIBA). J Virol Methods 2019;263:75-80. [Crossref] [PubMed]
- Aab A, Wirz O, van de Veen W, et al. Human rhinoviruses enter and induce proliferation of B lymphocytes. Allergy 2017;72:232-43. [Crossref] [PubMed]
- Feldman AS, He Y, Moore ML, et al. Toward primary prevention of asthma. Reviewing the evidence for early-life respiratory viral infections as modifiable risk factors to prevent childhood asthma. Am J Respir Crit Care Med 2015;191:34-44. [Crossref] [PubMed]
- Kotaniemi-Syrjänen A, Vainionpaa R, Reijonen TM, et al. Rhinovirus-induced wheezing in infancy--the first sign of childhood asthma? J Allergy Clin Immunol 2003;111:66-71. [Crossref] [PubMed]
- Jamieson KC, Warner SM, Richard L, et al. Rhinovirus in the Pathogenesis and Clinical Course of Asthma. Chest 2015;148:1508-16. [Crossref] [PubMed]
- Martínez-Roig A, Salvadó M, Caballero-Rabasco MA, et al. Viral coinfection in childhood respiratory tract infections. Arch Bronconeumol 2015;51:5-9. [Crossref] [PubMed]
- Henrickson KJ. Advances in the laboratory diagnosis of viral respiratory disease. Pediatr Infect Dis J 2004;23:S6-S10. [Crossref] [PubMed]
- Mogensen I, Alving K, Jacinto T, et al. Simultaneously elevated FeNO and blood eosinophils relate to asthma morbidity in asthmatics from NHANES 2007-12. Clin Exp Allergy 2018;48:935-43. [Crossref] [PubMed]
- Petsky HL, Cates CJ, Kew KM, et al. Tailoring asthma treatment on eosinophilic markers (exhaled nitric oxide or sputum eosinophils): a systematic review and meta-analysis. Thorax 2018;73:1110-9. [Crossref] [PubMed]
- Gadish T, Soferman R, Merimovitch T, et al. Exhaled nitric oxide in acute respiratory syncytial virus bronchiolitis. Arch Pediatr Adolesc Med 2010;164:727-31. [Crossref] [PubMed]
- Marguet C, Bocquel N, Benichou J, et al. Neutrophil but not eosinophil inflammation is related to the severity of a first acute epidemic bronchiolitis in young infants. Pediatr Allergy Immunol 2008;19:157-65. [Crossref] [PubMed]
- Huang Q, Deng X, Yan Z, et al. Establishment of a reverse genetics system for studying human bocavirus in human airway epithelia. Plos Pathog 2012;8:e1002899. [Crossref] [PubMed]
- Martinez FD. Development of wheezing disorders and asthma in preschool children. Pediatrics 2002;109:362-7. [PubMed]
- Djukanović R. Predicting Asthmatic Responses to Inhaled Allergen Using an Unbiased Transcriptomics Approach. Am J Respir Crit Care Med 2018;197:415-6. [Crossref] [PubMed]
- Hong JG. Improve continuously the management of childhood asthma. Zhonghua Er Ke Za Zhi 2008;46:721-3. [PubMed]
(English Language Editor: J. Chapnick)