Advances in genetic studies of inherited bone marrow failure syndromes and their associated malignancies
Introduction
The term bone marrow failure describes the functional loss of production of blood cells from hematopoietic stem cells. Studies demonstrated that there are more than 80 causative genes involved in bone marrow failure disorders (including acquired and inherited); but in about 40% of cases the cause is not known (1,2).
Inherited bone marrow failure syndromes (IBMFS) are complex mixture of genetic disorders characterised by insufficient blood cells production usually association with one or more somatic abnormality and an increased cancer risk.
There are more than 30 different types of disorders which are classified into IBMFS and the common types are: fanconi anemia (FA), dyskeratosis congenital (DC), shwachman-diamond syndrome (SDS), diamond-blackfan anemia (DBA), congenital amegakaryocytic thrombocytopenia (CAMT), severe congenital neutropenia (SCN) and thrombocytopenia absent radii (TAR) as summarised in Table 1 (3,4).
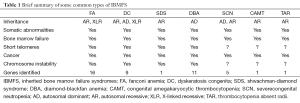
Full table
Recently more mutated genes causing IBMFS have been identified that show genomic instability, defects in DNA repair and telomere biology as genetic causes. Studies of disease pathophysiology at the genetic level have provided insights into several biological pathways, thereby correlation between genotype and phenotype, and clinical therapeutical strategies. This article reviews the latest studies of eight subtypes of the IBMFS and their associated malignancies.
Fanconi anemia
FA is usually inherited as an autosomal recessive (AR) trait, but it can also be X-linked recessive (XLR). FA is the most common and representative type of IBMFS. Patients with FA are characterised with progressive bone marrow failure, congenital abnormalities and susceptibility to malignancies, commonly in MDS, AML, and solid tumors, commonly as carcinoma of the oropharynx and skin (5). The first reported case of leukemia in patients with FA was in 1927 (6). Cells from FA patients exhibit a hypersensitivity to DNA interstrand cross-linking agents such as mitomycin and diepoxybutane. Patients with FA develop one or more types of cancers with the most common types being AML, MDS, head and neck squamous cell carcinoma, liver tumors, vaginal squamous cell carcinoma, and brain tumors. Recent studies found an association of FA with a pattern of recurrent chromosomal abnormalities including monosomy chromosome 7, deletion of the long arm of chromosome 7, gain of the long arms of chromosome 3 and 1 and the RUNX1 gene mutations in about 20% of the combined MDS cases. Patients with chromosome abnormalities involving 7 and 3 had a poor prognostic significance in disease (7).
The presence of specific types of genomic instability offers opportunity to classify the 16 subtypes/complementation groups in FA to elucidate the mechanism including the genes responsible. FA involved genes code for proteins involved in DNA repair. Eight (FANCA, FANCB, FANCC, FANCE, FANCF, FANCG, FANCL and FANCM) of the 16 FA proteins formed a FA core complex interacting with each other to active the FANCI-FANCD2 protein complex to a monoubiquitinated form to bind with DNA repair proteins (8,9). Four (FANCD1—also known as BRCA2, FANCJ, FANCN and FANCO) of the 16 FA genes are found to be associated with breast cancer (10). These findings suggest a link between FA and cancer through a faulty DNA repair mechanism.
Dyskeratosis congenita
DC or DKC is defined as one of the IBMFS characterised by the presence of bone marrow failure and the mucocutaneous triad of abnormal skin pigmentation, nail dystrophy, and mucosal leucoplakia (11). DC is an inherited disease with autosomal dominant (AD), AR and X-linked inheritance patterns. Patients with DC have increased risk of MDS, AML and other types of cancers (carcinomas of the upper gastro-intestinal tract). Aplastic anemia, MDS and AML in patient with DC is the first clinical sign. DC is the most typical representative type in IBMFS in telomere abnormality causing genomic instability due to the accelerated telomere shortening resulted in cell loss or dysfunction (12).
The nine genes responsible for DC (DKC1, TERT, TERC, TINF2, RTEL1, NOP10, NHP2, WRAP53 and CTC1) have been identified and responsible for functioning and maintenance of telomeres (2).
Shwachman-diamond syndrome
SDS is an AR disorder characterised by early onset exocrine pancreatic insufficiency, bone marrow failure and other genetic abnormalities. About 20% of SDS patients will have MDS and 25% patients will have leukemia (13). Deletions of chromosomal 7 and 20 are the most frequent abnormalities seen in patients with SDS presenting with MDS but such types of chromosomal abnormalities do not contribute to leukemia transformation. Molecular studies showed about 90% of SDS patients have SBDS gene mutation and its product has an important role in the maturation of the 60S ribosomal subunit (14).
Diamond-blackfan anemia
DBA is grouped with the early infant anemia and it is selectively seen in the erythroid lineage (pure red cell aplasia) with some somatic abnormalities such as craniofacial thumb, cardiac and urogenital malformations (15). An increased predisposition to MDS, AML and other types of tumors has been reported (16). In 1999, DBA gene (RPS19) was identified (17). Subsequently, heterozygous mutations in other encoding genes for ribosomal proteins of the small (RPS24, RPS17, RPS7, RPS10, RPS26) and large (RPL5, RPL11, RPL35) ribosomal subunits have also found to associated with DBA and RPS5 gene was found and is associated with patients with multiple physical abnormalities (18). RPS19 mutations causing DBA showed some ethnic difference in phenotype expression (19). Recent studies using a CGH identified deletion of RPL15 as a novel cause of DBA (20).
Severe congenital neutropenia
SCN is an AR disorder characterised by early age onset neutropenia and presented with recurrent life threatening infections with physical abnormalities. SCN can develop to MDS and AML with the secondary mutations including the granulocyte colony stimulating factor receptor gene (21). The neutrophil elastase gene (ELA2), defects in mitochondria gene (HAX1), deficiency in adenylate kinase 2 gene (AK2) and other 2 genes mutated (GFI1, WASP the transcriptional repressor and the cytoskeletal regulator respectively) associated with apoptosis were found to be responsible for SCN in at least 50% patients (22,23). Genetic defects in multiple pathways have been found to cause congenital neutropenia by controlling granulocytic progenitor differentiation.
Congeneital amegakaryocytic thrombocytopenia
CAMT is characterised by haemorrhages or bruises associated with thrombocytopenia in infancy but rarely presents with physical defects. MDS and AML but not solid tumors are associated with CAMT (24). Molecular studies demonstrate that the gene mutation called MPL (encoding of the receptor for thrombopoietin was responsible for CAMT and showed the correlation between genotype and phenotype (25).
Thrombocytopenia absent radii
TAR can be either an AR or de novo pattern, typically seen in infancy presenting with thrombocytopenia (low platelet count) and allergy to cow milk, physical characteristic of bilateral absent radii and other types of birth defects (26). Similarly, acute leukemia and solid tumors have been reported from patients with TAR (27).
The pathological mechanism of thrombocytopenia in TAR was studied and it was found that serum levels of thrombopoietin (the megakaryocyte growth factor) were increased suggesting an abnormal differentiation mechanism of megakaryocyte and platelet production (28).
The first molecular finding of interstitial microdeletion at chromosome 1q21.1 containing ten genes including the TAR responsible gene-RBM8A by using comparative genomic hybridisation (CGH) microarray technique was in 2007 (29). Recent findings show that RBM8A encodes the conserved Y14 subunit of the exon-junction complex that is essential for RNA processing and expressed in all hematopoietic lineages as the cause of TAR (30).
Conclusions
In the past our understanding of IBMFs was limited by several factors, such as, rarity of reported cases and the lack of DNA and other technologies. Traditionally, the diagnosis of IBMFS mainly relied on the uneven clinical features and limited laboratory tests. Even up to now, not all subtypes of IBMFs are well documented for the clinical and laboratory diagnostic criteria and guidelines due to heterogeneous complexity in diseases, particularly in the developing counties. About 45% of patients with IBMFS do not have mutations in known genes and further genetic identification is required.
The treatment strategy for IBMFS associated malignancies needs to be specifically tailored compared to other groups of patients with malignancies. Hence, the differential diagnosis on such malignancies such as MDS, acute leukemia and solid tumors is imperative.
The recent advances in molecular studies on the identifications of responsible genes/the defects of their pathways of IBMFS have provided more understandings in the pathophysiological mechanisms. Such advances also provided the link between IBMFS and some types of cancers via defective molecular pathways.
Next generation technology will make studies on IBMFS fast in the genetic etiology and the molecular pathways, clinical diagnosis and therapies so that there will be more IBMFS to be identified which currently remain unclassified. The studies on IBMFS offer a proxy for research in somatic cancer and in aging.
Acknowledgements
None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Parikh S, Bessler M. Recent insights into inherited bone marrow failure syndromes. Curr Opin Pediatr 2012;24:23-32. [PubMed]
- Tsangaris E, Klaassen R, Fernandez CV, et al. Genetic analysis of inherited bone marrow failure syndromes from one prospective, comprehensive and population-based cohort and identification of novel mutations. J Med Genet 2011;48:618-28. [PubMed]
- Dokal I, Vulliamy T. Inherited bone marrow failure syndromes. Haematologica 2010;95:1236-40. [PubMed]
- Alter BP, Giri N, Savage SA, et al. Malignancies and survival patterns in the National Cancer Institute inherited bone marrow failure syndromes cohort study. Br J Haematol 2010;150:179-88. [PubMed]
- Alter BP. Diagnosis, genetics, and management of inherited bone marrow failure syndromes. Hematology Am Soc Hematol Educ Program 2007.29-39. [PubMed]
- Fanconi G. Familiaere infantile perniziosaartige Anaemie (pernizioeses Blutbild und Konstitution). Jahrbuch Kinderheild 1927;117:257-80.
- Quentin S, Cuccuini W, Ceccaldi R, et al. Myelodysplasia and leukemia of Fanconi anemia are associated with a specific pattern of genomi cabnormalities that includes cryptic RUNX1/AML1 lesions. Blood 2011;117:e161-70. [PubMed]
- de Winter JP, Joenje H. The genetic and molecular basis of Fanconi anemia. Mutat Res 2009;668:11-9. [PubMed]
- Sakaguchi H, Nakanishi K, Kojima S. Inherited bone marrow failure syndromes in 2012. Int J Hematol 2013;97:20-9. [PubMed]
- Howlett NG, Taniguchi T, Olson S, et al. Biallelic inactivation of BRCA2 in Fanconi anemia. Science 2002;297:606-9. [PubMed]
- Walne AJ, Dokal I. Advances in the understanding of dyskeratosis congenita. Br J Haematol 2009;145:164-72. [PubMed]
- Vulliamy TJ, Knight SW, Mason PJ, et al. Very short telomeres in the peripheral blood of patients with X-linked and autosomal dyskeratosis congenita. Blood Cells Mol Dis 2001;27:353-7. [PubMed]
- Boocock GR, Morrison JA, Popovic M, et al. Mutations in SBDS are associated with Shwachman-Diamond syndrome. Nat Genet 2003;33:97-101. [PubMed]
- Wong CC, Traynor D, Basse N, et al. Defective ribosome assembly in Shwachman-Diamond syndrome. Blood 2011;118:4305-12. [PubMed]
- Ganapathi KA, Austin KM, Lee CS, et al. The human Shwachman-Diamond syndrome protein, SBDS, associates with ribosomal RNA. Blood 2007;110:1458-65. [PubMed]
- Shimamura A, Alter BP. Pathophysiology and management of inherited bone marrow failure syndromes. Blood Rev 2010;24:101-22. [PubMed]
- Gazda HT, Sheen MR, Vlachos A, et al. Ribosomal protein L5 and L11 mutations are associated with cleft palate and abnormal thumbs in Diamond-Blackfan anemia patients. Am J Hum Genet 2008;83:769-80. [PubMed]
- Vlachos A, Rosenberg PS, Atsidaftos E, et al. Incidence of neoplasia in Diamond Blackfan anemia: a report from the Diamond Blackfan Anemia Registry. Blood 2012;119:3815-9. [PubMed]
- Boultwood J, Pellagatti A, Wainscoat JS. Haploinsufficiency of ribosomal proteins and p53 activation in anemia: Diamond-Blackfan anemia and the 5q-syndrome. Adv Biol Regul 2012;52:196-203. [PubMed]
- Khincha PP, Savage SA. Genomic characterization of the inherited bone marrow failure syndromes. Semin Hematol 2013;50:333-47. [PubMed]
- Welte K, Zeidler C. Severe congenital neutropenia. Hematol Oncol Clin North Am 2009;23:307-20. [PubMed]
- Dale DC, Person RE, Bolyard AA, et al. Mutations in the gene encoding neutrophil elastase in congenital and cyclic neutropenia. Blood 2000;96:2317-22. [PubMed]
- Klein C. Genetic defects in severe congenital neutropenia: emerging insights into life and death of human neutrophil granulocytes. Annu Rev Immunol 2011;29:399-413. [PubMed]
- King S, Germeshausen M, Strauss G, et al. Congenital amegakaryocytic thrombocytopenia: a retrospective clinical analysis of 20 patients. Br J Haematol 2005;131:636-44. [PubMed]
- Ihara K, Ishii E, Eguchi M, et al. Identification of mutations in the c-mpl gene in congenital amegakaryocytic thrombocytopenia. Proc Natl Acad Sci U S A 1999;96:3132-6. [PubMed]
- Toriello HV. Thrombocytopenia-absent radius syndrome. Semin Thromb Hemost 2011;37:707-12. [PubMed]
- Albers CA, Paul DS, Schulze H, et al. Compound inheritance of a low-frequency regulatory SNP and a rare null mutation in exon-junction complex subunit RBM8A causes TAR syndrome. Nat Genet 2012;44:435-9, S1-2.
- Geddis AE. Congenital amegakaryocytic thrombocytopenia and thrombocytopenia with absent radii. Hematol Oncol Clin North Am 2009;23:321-31. [PubMed]
- Klopocki E, Schulze H, Strauss G, et al. Complex inheritance pattern resembling autosomal recessive inheritance involving a microdeletion in thrombocytopenia-absent radius syndrome. Am J Hum Genet 2007;80:232-40. [PubMed]
- Albers CA, Newbury-Ecob R, Ouwehand WH, et al. New insights into the genetic basis of TAR (thrombocytopenia-absent radii) syndrome. Curr Opin Genet Dev 2013;23:316-23. [PubMed]


